A Complete LAMP Protocol for SARS-CoV-2 Detection: Principles, Optimization, and Clinical Validation
This article provides a comprehensive guide for researchers and drug development professionals on implementing Loop-Mediated Isothermal Amplification (LAMP) for SARS-CoV-2 detection.
A Complete LAMP Protocol for SARS-CoV-2 Detection: Principles, Optimization, and Clinical Validation
Abstract
This article provides a comprehensive guide for researchers and drug development professionals on implementing Loop-Mediated Isothermal Amplification (LAMP) for SARS-CoV-2 detection. We cover the foundational principles of LAMP technology, including its mechanism and advantages over RT-PCR for point-of-care applications. A detailed, step-by-step methodological protocol is presented, from primer design and reaction setup to result interpretation. Critical troubleshooting and optimization strategies are discussed to enhance sensitivity, specificity, and robustness. Finally, the article examines validation frameworks, comparative performance metrics against gold-standard methods, and regulatory considerations. This guide serves as a practical resource for developing, optimizing, and validating robust LAMP assays for COVID-19 research and diagnostic development.
Understanding LAMP Technology: Principles and Advantages for SARS-CoV-2 Detection
Application Notes
Loop-mediated isothermal amplification (LAMP) is a rapid, specific, and efficient nucleic acid amplification technique. Its core mechanism for isothermal operation relies on the unique enzymatic properties of Bacillus stearothermophilus (Bst) DNA polymerase large fragment and a sophisticated primer design scheme. This enables autocycling strand displacement DNA synthesis at a constant temperature (60-65°C), eliminating the need for a thermal cycler.
Within the context of SARS-CoV-2 research, LAMP's isothermal nature makes it ideal for developing point-of-care diagnostics and high-throughput screening protocols. The use of Bst polymerase is central to this application, as it provides robust activity under isothermal conditions while maintaining high fidelity for the target sequences, such as the N, E, or ORF1ab genes of SARS-CoV-2.
Core Mechanistic Principles
- Strand Displacement Activity: Bst polymerase lacks 5'→3' exonuclease activity but possesses strong 5'→3' strand-displacing activity. This allows it to unwind downstream DNA without the need for thermal denaturation cycles, enabling isothermal amplification.
- Primer Design: A set of four to six primers (two inner and two outer, plus optional loop primers) recognize six to eight distinct regions on the target DNA. This ensures extremely high specificity.
- Formation of Loop Structures: The inner primers contain complementary sequences at their 5' ends. After initial extension, these sequences self-anneal to form looped DNA structures, which serve as initiation points for subsequent amplification rounds, leading to exponential synthesis.
- Amplification Products: The reaction yields a mixture of stem-loop DNAs with various lengths and cauliflower-like structures with multiple inverted repeats.
Key Quantitative Data on Bst Polymerase in LAMP
Table 1: Characteristics of Bst DNA Polymerase Large Fragment
| Property | Typical Value/Range | Significance for LAMP |
|---|---|---|
| Optimal Temperature | 60 - 65°C | Enables single-temperature incubation. |
| Processivity | High | Synthesizes long DNA fragments without dissociating, accelerating amplification. |
| Strand Displacement | Active | Eliminates need for thermal denaturation; core to isothermal mechanism. |
| 5'→3' Exonuclease | Inactive | Prevents undesired degradation of primers and loop structures. |
| Reverse Transcriptase | Available in variants (Bst 2.0/3.0) | Enables RT-LAMP for RNA viruses like SARS-CoV-2 in a one-step protocol. |
| Half-life | >2 hours at 65°C | Supports long, robust reactions for high sensitivity. |
| Mg²⁺ Requirement | 4-8 mM | Often optimized in commercial buffers; crucial for polymerase activity. |
| dNTP Consumption | High (~millimolar) | Due to high yield; requires sufficient concentration in master mix. |
Table 2: Comparison of LAMP Performance with Bst Polymerase for SARS-CoV-2 Detection
| Parameter | Typical Performance Range | Notes |
|---|---|---|
| Amplification Time | 15 - 60 minutes | Depends on target copy number, primer set, and detection method. |
| Limit of Detection (LoD) | 10 - 100 copies/reaction | Comparable to, and sometimes surpassing, conventional PCR. |
| Specificity | Very High | Due to multi-primer recognition sites; must be validated in silico and empirically. |
| Amplification Efficiency | Very High (10⁹ - 10¹⁰ copies in 1h) | Results in high yield, enabling visual detection (turbidity, color change). |
| Optimal Reaction Temperature | 62 - 65°C | Standard for DNA targets; ~60-63°C for one-step RT-LAMP. |
Experimental Protocols
Protocol 1: Standard RT-LAMP for SARS-CoV-2 RNA Detection
Objective: To detect SARS-CoV-2 RNA using a one-step reverse transcription LAMP (RT-LAMP) assay with Bst polymerase.
Research Reagent Solutions & Materials:
| Item | Function/Brief Explanation |
|---|---|
| Bst 2.0 or 3.0 WarmStart DNA Polymerase | Engineered Bst variant with high strand displacement and mesophilic reverse transcriptase activity for one-step RT-LAMP. |
| LAMP Primer Mix (F3/B3, FIP/BIP, LF/LB) | Specifically designed for SARS-CoV-2 target (e.g., N gene). Provides specificity and enables loop formation. |
| WarmStart Colorimetric LAMP 2X Master Mix | Commercial mix containing dNTPs, buffer, MgSO4, and a pH-sensitive dye for visual readout. |
| RNA Template | Extracted viral RNA from nasopharyngeal swabs or synthetic control. |
| Nuclease-free Water | To bring reaction to volume without degrading primers/template. |
| Positive & Negative Controls | SARS-CoV-2 RNA and nuclease-free water, essential for validation. |
| Heating Block or Water Bath | Maintains constant isothermal temperature (e.g., 63°C). |
Methodology:
- Primer Design/Selection: Use validated primer sets for SARS-CoV-2 (e.g., from Zhang et al., J Clin Microbiol, 2020). Resuspend primers to stock concentrations (e.g., 100 µM for F3/B3, 40 µM for Loop primers). Prepare a working primer mix.
- Reaction Setup (25 µL total volume):
- 12.5 µL 2X Colorimetric LAMP Master Mix
- 5 - 7.5 µL Primer Mix (final concentrations: 0.2 µM F3/B3, 1.6 µM FIP/BIP, 0.8 µM LF/LB)
- 2 - 5 µL RNA template (or control)
- Nuclease-free water to 25 µL
- Amplification:
- Incubate reactions at 63°C for 30-45 minutes in a heating block or water bath.
- No initial denaturation or final extension is required.
- Result Interpretation (Colorimetric):
- Positive: Color changes from pink to yellow due to acidification of the reaction (pyruvate production).
- Negative: Remains pink.
- Include a no-template control (NTC) and positive control in every run.
Protocol 2: Turbidity-Based Real-Time Monitoring of LAMP
Objective: To monitor LAMP amplification kinetics in real-time via magnesium pyrophosphate precipitate formation.
Research Reagent Solutions & Materials:
| Item | Function/Brief Explanation |
|---|---|
| Bst DNA Polymerase, Large Fragment | Standard strand-displacing polymerase. |
| LAMP Primer Mix | As in Protocol 1. |
| Isothermal Amplification Buffer | Contains Tris-HCl, (NH4)2SO4, KCl, MgSO4, Tween 20. |
| Betaine (5M stock) | Additive that destabilizes DNA secondary structures, improving primer annealing and efficiency. |
| dNTP Solution | Deoxynucleotide triphosphates, energy source and building blocks. |
| Calcein/Mn²⁺ Dye System (Optional) | Fluorometric indicator for real-time fluorescence detection. |
| Real-time Isothermal Fluorometer or qPCR Thermocycler | Equipment capable of maintaining constant temperature and taking periodic absorbance (600 nm) or fluorescence readings. |
Methodology:
- Reaction Setup (25 µL):
- 1X Isothermal Amplification Buffer (with 6-8 mM MgSO4)
- 1.4 mM each dNTP
- 0.8 M Betaine
- Primer concentrations as in Protocol 1
- 8 U Bst DNA polymerase
- Template DNA/RNA
- (For fluorescence) 25 µM Calcein, 0.5 mM MnCl2
- Amplification & Monitoring:
- Place tubes/strips in instrument preheated to 65°C.
- Monitor absorbance at 600 nm every 30-60 seconds for 60 minutes.
- Turbidity increases as magnesium pyrophosphate precipitates.
- Data Analysis:
- Plot time (x-axis) vs. absorbance (y-axis).
- The time to reach a threshold absorbance (Tt) is inversely proportional to the initial template concentration, enabling quantitative analysis.
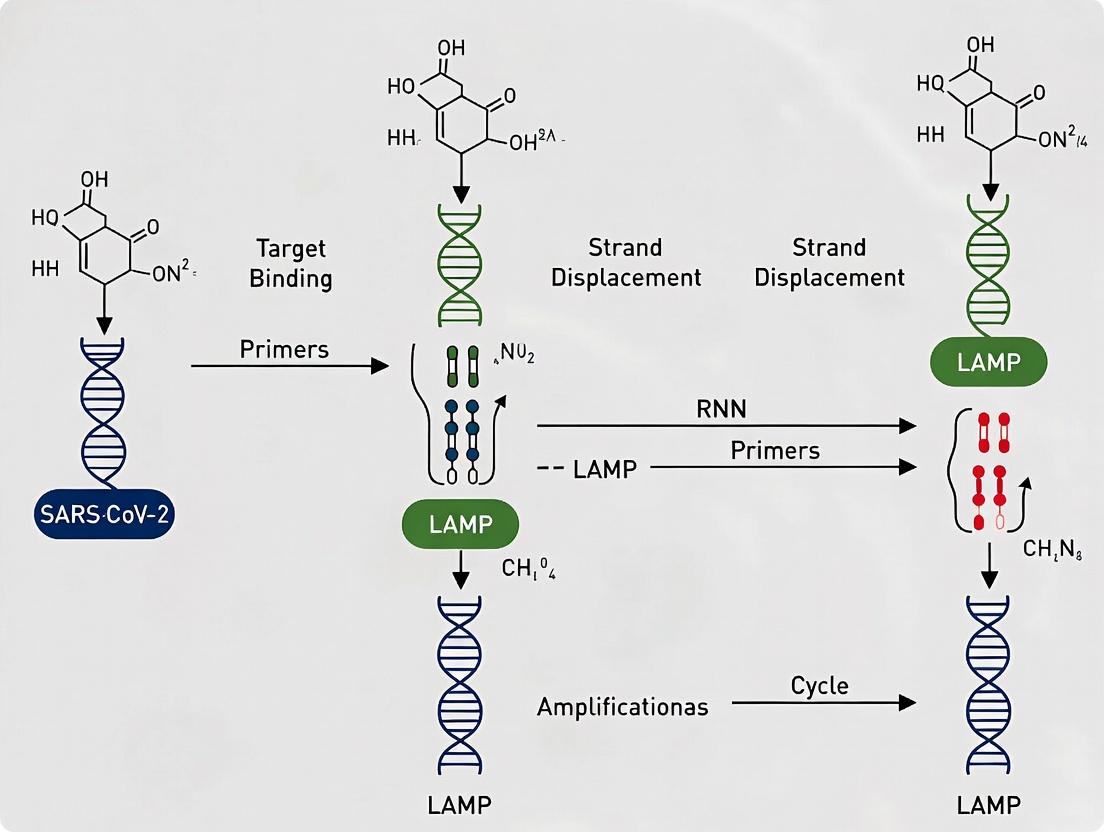
Mechanistic & Workflow Diagrams
Title: LAMP Amplification Cycle with Bst Polymerase
Title: SARS-CoV-2 RT-LAMP Experimental Workflow
Within the broader context of developing a robust, field-deployable LAMP protocol for SARS-CoV-2 detection, the precise formulation of reaction components is critical. This document details the application notes and protocols for three foundational elements: primers, buffer, and visual indicators, focusing on their optimization for sensitive and specific viral RNA detection.
Primers: Design and Specificity
LAMP employs six primers targeting eight distinct regions on the target DNA. For SARS-CoV-2, primers are typically designed against conserved regions such as the N (nucleocapsid), E (envelope), or ORF1ab genes.
Key Quantitative Data: Primer Sequences and Concentrations Table 1: Typical Primer Set for SARS-CoV-2 N Gene Detection
| Primer Name | Type | Sequence (5' -> 3') | Final Concentration (μM) | Function |
|---|---|---|---|---|
| F3 | Forward Outer | ACGCCGTAACGGCACCAAG | 0.2 | Initiates strand displacement |
| B3 | Backward Outer | CAGTGCTGGTTCACACCTTGTC | 0.2 | Initiates strand displacement |
| FIP (F1c+F2) | Forward Inner | TCTGGTTACTGCCAGTTGAATCTGGAAGAGACAGTTGC | 1.6 | Forms loop structure; main amplification primer |
| BIP (B1c+B2) | Backward Inner | CAGGCATGGCAAACAACTCAGCAACACTATTAGCAATG | 1.6 | Forms loop structure; main amplification primer |
| LF | Loop Forward | GCAGCAGTAGGCAAGCACTT | 0.8 | Accelerates amplification by binding loop |
| LB | Loop Backward | CTGGTAGGCTTGAAGTGTCG | 0.8 | Accelerates amplification by binding loop |
Protocol 1: Primer Design and Validation Workflow
- Target Selection: Align sequences of SARS-CoV-2 variants (e.g., from GISAID) to identify highly conserved ~200 bp region.
- Design: Use software (e.g., PrimerExplorer V5) to generate candidate primer sets.
- Specificity Check: Perform in silico PCR against human genome and common respiratory flora databases.
- Empirical Testing: Test primer sets against synthetic SARS-CoV-2 RNA (10^2 to 10^6 copies/μL) and negative controls (no template, human genomic DNA). Evaluate time-to-positive (Tp) and amplification efficiency.
Title: Primer Design and Validation Workflow for LAMP
Reaction Buffer: Optimizing the Biochemical Environment
The buffer sustains isothermal amplification by providing optimal pH, salt conditions, and co-factors for the Bst DNA polymerase.
Key Quantitative Data: Standard LAMP Buffer Composition Table 2: Composition of a Standard 2x LAMP Reaction Buffer
| Component | Final Concentration | Function & Rationale |
|---|---|---|
| Tris-HCl (pH 8.8) | 40 mM | Maintains optimal pH for Bst polymerase activity. |
| KCl | 50 mM | Salt concentration crucial for primer annealing and polymerase processivity. |
| MgSO₄ | 8-10 mM | Critical co-factor for Bst polymerase. Excess can lead to non-specific amplification. |
| Betaine | 0.8-1.2 M | Reduces secondary structure in DNA, improving primer access and strand displacement. |
| dNTPs | 1.4 mM each | Nucleotide building blocks for DNA synthesis. |
| Tween-20 | 0.2% (v/v) | Stabilizes polymerase and reduces surface adsorption. |
| Bst 2.0/3.0 Polymerase | 8-16 U/reaction | Engineered for high strand displacement activity at 60-65°C. |
Protocol 2: Mg²⁺ and Betaine Concentration Optimization
- Prepare a master mix containing all components except MgSO₄ and betaine.
- Aliquot the master mix into 8 tubes.
- Create a matrix of MgSO₄ (6, 8, 10, 12 mM final) and betaine (0.6, 1.0 M final) concentrations.
- Add synthetic SARS-CoV-2 RNA target (10^3 copies/reaction) to each tube.
- Incubate at 63°C for 45 minutes.
- Measure Tp using a real-time turbidimeter or fluorescent dye. The condition with the lowest Tp and no false-positive in NTC is optimal.
Visual Indicators: Enabling Endpoint Detection
Visual indicators allow result interpretation without instrumentation, crucial for point-of-care applications.
Key Quantitative Data: Common Visual Indicators Table 3: Comparison of Visual Detection Methods for SARS-CoV-2 LAMP
| Indicator Type | Mechanism | Typical Result (Positive/Negative) | Time to Read | Notes |
|---|---|---|---|---|
| Colorimetric (pH) | Proton release during amplification lowers pH. | Yellow -> Magenta (or Orange) / Remains Yellow | Post-amplification (~30-45 min) | Requires optimized buffer with phenol red; prone to condenser contamination artifacts. |
| Fluorescent (Intercalating Dye) | Dye (e.g., SYBR Green I) binds dsDNA and fluoresces under UV/blue light. | Bright Green / Remains Orange | Post-amplification or real-time | Caution: SYBR Green I is a potent mutagen; add post-amplension or use closed-tube variants. |
| Pyrophosphate (Turbidity) | Magnesium pyrophosphate precipitate formation. | Turbid / Clear | Real-time or endpoint | Measured optically at 400 nm; compatible with real-time monitoring. |
| Metal Ion Sensors | Calcein quenching by Mn²⁺ is reversed by pyrophosphate. | Green Fluorescence / Orange Quenched | Post-amplification | Pre-formulated kits often use this system. |
Protocol 3: Closed-Tube Detection with Hydroxy Naphthol Blue (HNB)
- Reaction Setup: Prepare standard LAMP master mix (as optimized in Protocol 2). Add HNB to a final concentration of 120 μM before amplification.
- Initial Color: The reaction mix will be violet due to the HNB-Mg²⁺ complex.
- Amplification: Run the reaction at 63°C for 45 minutes.
- Endpoint Reading: Positive amplification consumes Mg²⁺ (into magnesium pyrophosphate), causing a color shift from violet to sky blue. Negative reactions remain violet. Do not open tubes.
Title: Mechanism of HNB Colorimetric Detection in LAMP
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Materials for SARS-CoV-2 LAMP Protocol Development
| Item | Function | Example/Supplier (Research-Use Only) |
|---|---|---|
| Bst 2.0/3.0 WarmStart Polymerase | High-activity, thermostable DNA polymerase with strand displacement. | New England Biolabs, Thermo Fisher Scientific. |
| Synthetic SARS-CoV-2 RNA Control | Quantitative positive control for assay development and validation. | Twist Biosciences, ATCC. |
| Human Genomic DNA & Respiratory Panel RNA | Negative controls to assess specificity and cross-reactivity. | Promega, ZeptoMetrix. |
| 2x LAMP Master Mix (Colorimetric) | Pre-optimized buffer, salts, dNTPs, and indicator (e.g., phenol red). | Lucigen, OptiGene. |
| RNase Inhibitor | Protects viral RNA template during reverse transcription-LAMP (RT-LAMP) setup. | Takara, PrimeSafe. |
| Rapid Heat Block or Dry Bath | Provides precise isothermal incubation at 60-65°C. | ThermoFisher, Major Science. |
| Portable Fluorimeter or Turbidimeter | For real-time, quantitative monitoring of LAMP reactions. | Genie HT (OptiGene), LA-500 (Eiken). |
Why LAMP for SARS-CoV-2? Advantages in Speed, Simplicity, and Point-of-Care Potential.
Within the context of a broader thesis on LAMP for SARS-CoV-2, this application note details the rationale and methodologies underpinning its adoption. Loop-mediated isothermal amplification (LAMP) has emerged as a critical molecular tool for the detection of SARS-CoV-2 RNA, offering distinct advantages over traditional RT-qPCR, particularly in decentralized testing scenarios. Its principal benefits lie in operational speed, technical simplicity, and inherent compatibility with point-of-care (POC) device integration.
Quantitative Advantages: LAMP vs. RT-qPCR
The following table summarizes key performance and operational metrics.
Table 1: Comparative Analysis of SARS-CoV-2 Detection Methods
| Parameter | RT-qPCR | LAMP | Implication for LAMP |
|---|---|---|---|
| Amplification Temperature | Thermo-cycling (55-60°C denaturation, ~60°C annealing/extension) | Isothermal (60-65°C constant) | Eliminates need for precise thermal cycler; enables use of simple heat blocks. |
| Time-to-Result | 60 - 120 minutes | 15 - 45 minutes | Faster result generation critical for screening and rapid decision-making. |
| Instrument Complexity | High (precise thermal cycling, fluorescence detection) | Low (constant temperature, visual detection possible) | Reduces cost and footprint; enables field deployment. |
| Detection Method | Fluorometric (probes or intercalating dyes) | Colorimetric (pH indicators, metal ion indicators), Fluorometric, or Turbidimetric | Visual readout eliminates need for detectors, simplifying the workflow. |
| Sensitivity | High (often < 100 copies/reaction) | Comparable/High (typically 10 - 1000 copies/reaction) | Suitable for clinical diagnosis given adequate viral loads. |
| Specificity | High (dependent on probe/primer design) | Very High (4-6 primers targeting 6-8 distinct regions) | Reduced false positives due to multiple primer recognition sites. |
| RNA Extraction Necessity | Typically required | Can be bypassed (with validated protocols) | Enables direct testing from swab samples, dramatically simplifying workflow. |
Detailed Protocol: Colorimetric RT-LAMP for SARS-CoV-2
This protocol details a one-step, colorimetric RT-LAMP assay suitable for research and potential POC development.
I. Principle: Reverse transcription and LAMP amplification occur concurrently in a single tube at 65°C. Amplification produces pyrophosphate ions, which bind magnesium ions in solution, reducing free Mg²⁺. This causes a pH-sensitive dye (e.g., phenol red) to shift from pink/red (alkaline) to yellow (acidic), providing a visual result.
II. Research Reagent Solutions & Essential Materials Table 2: The Scientist's Toolkit for SARS-CoV-2 RT-LAMP
| Item | Function | Example/Notes |
|---|---|---|
| LAMP Primer Mix | Contains 6 primers (F3, B3, FIP, BIP, LF, LB) targeting conserved regions of SARS-CoV-2 (e.g., N, E, Orf1ab genes). | Custom-designed, pre-mixed for stability. Critical for specificity. |
| Isothermal Master Mix | Provides buffer, dNTPs, MgSO4, and Bst DNA Polymerase (or Bst 2.0/WarmStart) with reverse transcriptase. | Commercial mixes (e.g., WarmStart LAMP Kit) ensure reproducibility. |
| pH-Sensitive Dye | Visual indicator of amplification (proton release). | Phenol Red (0.2 mM final) or Hydroxy Naphthol Blue. |
| RNA Template | Sample input. Can be purified RNA or inactivated viral transport media. | For direct assays, optimize sample volume (<25% of reaction). |
| Positive Control Template | Synthetic RNA or quantified viral RNA with target sequences. | Essential for validating each run. |
| Negative Control | Nuclease-free water and/or human RNA. | Controls for contamination and false positives. |
| Heating Block / Dry Bath | Maintains constant 60-65°C. | Simple, inexpensive equipment. POC devices use integrated heaters. |
| Microcentrifuge Tubes or Strip Tubes | Reaction vessels. | Preferably 0.2 mL tubes for rapid thermal equilibration. |
III. Step-by-Step Procedure:
- Reaction Setup (on ice or cold block):
- Prepare a master mix for n+2 reactions to account for pipetting loss.
- For a single 25 µL reaction, combine:
- 12.5 µL 2x Isothermal Master Mix with RTase
- 5 µL Primer Mix (final concentration: 1.6 µM FIP/BIP, 0.2 µM F3/B3, 0.4 µM LF/LB)
- 1 µL Phenol Red (0.2 mM stock)
- X µL RNA template (e.g., 5 µL)
- Nuclease-free water to 25 µL.
- Mix and Load:
- Mix gently by pipetting. Briefly centrifuge to collect contents.
- Aliquot 25 µL into individual reaction tubes.
- Amplification:
- Place tubes in a pre-heated heating block or dry bath at 65°C.
- Incubate for 30 minutes.
- Result Interpretation:
- Visual Assessment: A color change from pink/red to yellow indicates a positive result. No color change (remains pink/red) indicates a negative result.
- Quantitative Option (if using real-time fluorometer): Monitor fluorescence (intercalating dye) in real-time for Tt (time-threshold) calculation.
Visual Workflow and Logical Framework
Title: SARS-CoV-2 LAMP Testing Workflow
Title: Method Selection Decision Logic
Within the context of advancing LAMP (Loop-Mediated Isothermal Amplification) protocols for SARS-CoV-2 detection, the strategic selection of conserved genomic targets is paramount for assay robustness, especially against evolving variants. This application note details the methodology for identifying and validating highly conserved regions within the SARS-CoV-2 genome—specifically the Nucleocapsid (N), Envelope (E), and open reading frame 1ab (Orf1ab) genes—to ensure broad detection capability and diagnostic reliability.
The high mutation rate of RNA viruses like SARS-CoV-2 necessitates targeting evolutionarily stable genomic regions for diagnostic assays. LAMP’s isothermal amplification is highly sensitive but requires careful primer design to avoid mismatches with variant sequences. Conserved regions within structural (N, E) and replicase (Orf1ab) genes offer optimal targets for durable assay design.
Comparative Conservation Analysis of SARS-CoV-2 Genomic Regions
A bioinformatic analysis of publicly available SARS-CoV-2 sequences (GISAID, NCBI) was performed to quantify nucleotide conservation across key genes. The metric represents the percentage of sequences without mutations at each position, averaged across the gene region.
Table 1: Conservation Metrics for Key SARS-CoV-2 Genomic Targets
| Genomic Target | Region Length (nt) | Avg. Nucleotide Conservation (%)* | Key Variant Cross-Reactivity (Tested) | Suitability for LAMP Primer Design |
|---|---|---|---|---|
| N Gene | ~1260 | 99.2 | Alpha, Delta, Omicron (BA.1, BA.2, BA.5) | High (multiple conserved stretches) |
| E Gene | ~228 | 99.5 | All major VOCs | Moderate (shorter length) |
| Orf1ab (RdRp) | ~1323 | 98.8 | All major VOCs | High (long, highly conserved core) |
| S Gene | ~3822 | 96.1 | Limited (many mutational hotspots) | Low |
*Data derived from analysis of >1.5 million sequences (last 6 months).
Protocol: Identification and Validation of Conserved Regions for LAMP Assay Design
Bioinformatic Pipeline for Conservation Scoring
Objective: To identify regions of high sequence conservation within target genes for LAMP primer design (F3/B3, FIP/BIP, LF/LB).
Materials & Reagents:
- Sequence Database: GISAID EpiCoV database (requires registration and data access agreement).
- Software Tools: MAFFT (multiple sequence alignment), Geneious Prime or Biopython for conservation analysis, PrimerExplorer (Fujitsu) or LAMP Designer (Thermo Fisher) for primer design.
- Computing Resource: Local server or high-performance computing cluster for large-scale alignments.
Procedure:
- Data Curation: Download a representative set of SARS-CoV-2 complete genome sequences (minimum 5,000, spanning the pandemic timeline and major lineages).
- Sequence Alignment: Use MAFFT v7.475 with the
--autoflag to perform multiple sequence alignment against the reference genome (e.g., NC_045512.2). - Conservation Plotting: Calculate per-site nucleotide entropy or percent identity using a custom script (Python/Biopython) or alignment viewer. Identify regions >50 base pairs with conservation >99%.
- Primer Design: Input selected conserved regions into LAMP-specific primer design software. Set parameters: amplicon size 150-250 bp, Tm 58-65°C (inner primers), GC content 40-65%. Design 2-3 primer sets per target region.
- In Silico Specificity Check: Perform BLASTn analysis against the human genome and common respiratory flora to ensure specificity.
Experimental Validation of Conserved Target LAMP Assays
Objective: To validate the sensitivity and cross-reactivity of LAMP assays designed against conserved regions.
Research Reagent Solutions: Table 2: Essential Reagents for LAMP Validation
| Reagent / Material | Function in Protocol | Example Product / Note |
|---|---|---|
| Bst 2.0/3.0 DNA Polymerase | Isothermal amplification enzyme with strand displacement activity. | New England Biolabs Bst 2.0 WarmStart |
| Fluorescent DNA Intercalating Dye | Real-time monitoring of amplification. | SYTO 9, EvaGreen |
| Synthetic SARS-CoV-2 RNA Controls | Positive control for assay calibration and limit of detection (LoD) studies. | BEI Resources, ATCC VR-1986HK |
| RNase Inhibitor | Protects RNA template during reaction setup. | Recombinant RNasin |
| WarmStart RTx Reverse Transcriptase | For reverse transcription in one-step RT-LAMP. | New England Biolabs |
| Heat Block or Portable Dry Bath | Provides constant 60-65°C isothermal conditions. |
Procedure:
- One-Step RT-LAMP Reaction Setup (25 µL):
- 1x Isothermal Amplification Buffer
- 6 mM MgSO4
- 1.4 mM each dNTP
- 8 U Bst 2.0 WarmStart Polymerase
- 0.2 U WarmStart RTx Reverse Transcriptase
- 20 U RNase Inhibitor
- 1x Fluorescent Dye (e.g., 0.5x SYTO 9)
- Primer Mix: 1.6 µM FIP/BIP, 0.2 µM F3/B3, 0.4 µM LF/LB.
- 5 µL of template RNA (or nuclease-free water for NTC).
- Amplification Protocol:
- Incubate at 60°C for 30-45 minutes in a real-time fluorometer or qPCR instrument acquiring fluorescence every 30 seconds.
- Use a positive control (10^3 copies/µL synthetic RNA) and no-template control (NTC).
- Limit of Detection (LoD) Determination:
- Perform a 10-fold serial dilution of synthetic RNA (from 10^6 to 10^0 copies/µL).
- Run each dilution in replicates (n=8-12). The LoD is the lowest concentration detected in ≥95% of replicates.
- Cross-Reactivity (Specificity) Testing:
- Test the assay against RNA/DNA from common respiratory pathogens (e.g., influenza A/B, RSV, HCoV-229E, human genomic DNA) and SARS-CoV-2 variants of concern (VOC) RNA if available.
Visualizing the Experimental and Bioinformatics Workflow
Title: Workflow for Conserved Target LAMP Assay Development
Title: Key Conserved Genomic Targets for SARS-CoV-2 LAMP
Application Notes: The Journey of LAMP Technology
Loop-mediated isothermal amplification (LAMP) was first described by Notomi et al. in 2000 as a novel nucleic acid amplification method. Its core innovation was the use of a DNA polymerase with high strand displacement activity and a set of four to six primers that recognize six to eight distinct regions on the target DNA, enabling amplification under isothermal conditions (60–65°C). This eliminated the need for thermal cycling equipment, a key limitation of PCR.
The advent of the COVID-19 pandemic in late 2019 created an urgent, global demand for rapid, scalable, and field-deployable molecular diagnostics. LAMP's inherent advantages—speed (results in 15-60 minutes), robustness to inhibitors, and compatibility with simple heating blocks—made it an ideal candidate for SARS-CoV-2 detection. The technology evolved rapidly from a laboratory technique to a cornerstone of pandemic response, with numerous protocols developed for point-of-care testing, home testing, and wastewater surveillance.
Key milestones in its application for SARS-CoV-2 include the early design of primer sets targeting the ORF1ab, N, S, and E genes, the integration with colorimetric (pH-sensitive dyes) or fluorescent readouts for visual interpretation, and the development of lyophilized, room-stable reagents to enhance deployability.
Table 1: Comparative Performance of Representative SARS-CoV-2 LAMP Assays
| Assay Name/Target | Time to Result | Limit of Detection (LoD) | Sensitivity (%) | Specificity (%) | Key Feature |
|---|---|---|---|---|---|
| CDC N1/N2 LAMP | 40 min | ~100 copies/µL | 97.5 | 100 | Uses standard fluorophores |
| Colorimetric RT-LAMP (N gene) | 30 min | ~200 copies/reaction | 99.0 | 98.5 | Phenol red visual readout |
| SHERLOCK-based DETECTR | 45 min | 70 copies/µL | 95.0 | 100 | CRISPR-Cas12a coupled |
| Direct RT-LAMP (Saliva) | 35 min | 500 copies/mL | 94.2 | 97.1 | No RNA extraction step |
| Lyophilized RT-LAMP | 60 min | ~1000 copies/reaction | 91.7 | 100 | Room-temperature stable |
Detailed Experimental Protocols
Protocol 2.1: Two-Step Colorimetric RT-LAMP for SARS-CoV-2 RNA
Objective: To detect SARS-CoV-2 viral RNA using a reverse transcription (RT) and LAMP amplification with a visual colorimetric readout.
Key Research Reagent Solutions:
- WarmStart Colorimetric LAMP 2X Master Mix: Contains Bst 2.0/3.0 DNA polymerase, reverse transcriptase, pH-sensitive dye, dNTPs, and optimized buffer.
- SARS-CoV-2 Primers (N gene): A set of six primers (F3, B3, FIP, BIP, LF, LB) designed against a conserved region of the nucleocapsid gene.
- Nuclease-free Water: For dilution and negative controls.
- Positive Control Template: Synthetic RNA spanning the N gene target region.
- Sample Preparation Reagent (e.g., Proteinase K, Chelex-100): For crude sample preparation from nasal swabs or saliva.
Procedure:
- Primer Reconstitution: Resuspend lyophilized primer mix in nuclease-free water to create a 10X primer stock solution.
- Reaction Assembly (25 µL total volume):
- 12.5 µL WarmStart Colorimetric LAMP 2X Master Mix
- 2.5 µL 10X Primer Mix
- 5–10 µL of extracted RNA or processed sample (containing up to 10^6 copies of viral RNA)
- Add nuclease-free water to 25 µL.
- Controls: Always include a no-template control (NTC) with water and a positive control with synthetic RNA.
- Amplification: Incubate reactions in a heat block or dry bath at 65°C for 30-40 minutes. Do not use a thermocycler with a heated lid.
- Result Interpretation:
- Positive: Solution turns from pink to yellow due to acidification (proton release during amplification).
- Negative: Solution remains pink.
- Invalid: If the NTC turns yellow, indicates contamination.
Protocol 2.2: Direct Saliva RT-LAMP Protocol Without RNA Extraction
Objective: To enable rapid testing by bypassing the RNA extraction step, using heat-inactivated saliva samples.
Procedure:
- Sample Inactivation: Mix 100 µL of fresh saliva with 100 µL of DNA/RNA Shield or a proteinase K solution. Heat at 95°C for 5 minutes.
- Cooling: Centrifuge briefly and let cool to room temperature.
- Reaction Assembly (20 µL total):
- 10 µL of 2X LAMP Master Mix (with reverse transcriptase)
- 2 µL of 10X primer set
- 3 µL of inactivated saliva supernatant
- 5 µL nuclease-free water.
- Amplification & Detection: Incubate at 63°C for 45 minutes. Use a portable fluorimeter for real-time monitoring or visual inspection with SYBR Green I added post-amplification.
Visualizations
Title: SARS-CoV-2 LAMP Testing Workflow
Title: LAMP Primer Design and Binding Mechanism
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for SARS-CoV-2 LAMP Research
| Item | Function/Description | Example Product/Brand |
|---|---|---|
| Bst 2.0/3.0 DNA Polymerase | Engineered DNA polymerase with high strand displacement activity, essential for isothermal amplification. | New England Biolabs WarmStart Bst 2.0/3.0 |
| Reverse Transcriptase | For converting viral RNA to cDNA in one-step RT-LAMP protocols. | WarmStart RTx |
| LAMP Primer Sets (6 primers) | Specifically designed to recognize 8 regions of the SARS-CoV-2 genome. Critical for specificity and efficiency. | Custom synthesized (e.g., IDT, Metabion) |
| Colorimetric Master Mix | 2X mix containing Bst polymerase, buffers, dNTPs, and a pH indicator for visual readout. | WarmStart Colorimetric LAMP 2X Master Mix |
| Fluorescent DNA Intercalating Dye | Binds double-stranded DNA products for real-time or end-point fluorescence detection. | SYBR Green I, EvaGreen |
| Synthetic SARS-CoV-2 RNA Control | Quantified, non-infectious RNA for assay validation, standard curves, and positive controls. | Twist Synthetic SARS-CoV-2 RNA Control |
| Heat Block/Dry Bath | Provides consistent isothermal incubation at 60-65°C. Essential for field use. | ThermoFisher Digital Dry Bath |
| Rapid Extraction Kit | Simple, column-free RNA extraction reagents for fast sample prep. | MagMAX Viral/Pathogen Kit |
| Lyophilization Stabilizer | For creating room-temperature stable, single-reaction pellets for point-of-care use. | Trehalose, Pullulan |
Step-by-Step SARS-CoV-2 LAMP Protocol: From Primer Design to Data Analysis
Within the broader thesis on optimizing LAMP (Loop-Mediated Isothermal Amplification) for SARS-CoV-2 detection, the design of primers is the most critical determinant of assay success. LAMP employs four to six primers recognizing six to eight distinct regions on the target DNA, making strategic design paramount for specificity and amplification efficiency. This Application Note details the criteria and protocols for designing and validating LAMP primers for SARS-CoV-2 genomic targets.
Core Criteria for LAMP Primer Design
LAMP primer design extends beyond conventional PCR requirements. The following quantitative criteria, derived from current literature and bioinformatics guidelines, must be met.
Table 1: Quantitative Design Criteria for SARS-CoV-2 LAMP Primers
| Primer Type | Length (nt) | GC Content (%) | Tm Range (°C) | ΔG (3' end) | Specificity Check |
|---|---|---|---|---|---|
| F3 / B3 | 18-22 | 40-60 | 55-60 | > -9 kcal/mol | BLAST against human/human microbiome & SARS-CoV-2 variants |
| FIP / BIP | 40-45 total | 50-60 | 60-65 (overall) | 3' end: > -4 kcal/mol | Internal stability; avoid dimerization |
| LF / LB* | 18-22 | 40-55 | 60-65 | > -9 kcal/mol | Enhances speed; not always required |
*Loop primers (LF, LB) are designed between F1/F2 and B1/B2 regions to accelerate amplification.
Specificity Imperative: For SARS-CoV-2, primers must target conserved regions among variants (e.g., within N, E, or ORF1ab genes) while avoiding homology with the human genome and common respiratory tract flora.
Protocol: In Silico Design and Validation Workflow
Protocol 3.1: Primer Design Using PrimerExplorer or Alternative Software
Objective: To generate candidate primer sets for a defined SARS-CoV-2 target sequence. Materials: Target genome (e.g., NC_045512.2), primer design software (PrimerExplorer V5, NEB LAMP Designer), standard computer. Method:
- Sequence Selection: Identify a conserved 200-300 bp region from the SARS-CoV-2 genome (e.g., N gene).
- Software Input: Submit the FASTA sequence to PrimerExplorer V5. Set parameters per Table 1.
- Set Generation: The software outputs multiple candidate sets (F3, B3, FIP, BIP, LF, LB). Select 2-3 candidate sets for further analysis.
- Specificity Verification: Perform in silico PCR and BLAST each individual primer sequence against:
- The human reference genome (hg38).
- A database of common respiratory pathogens.
- A comprehensive SARS-CoV-2 variant database (GISAID).
- Secondary Structure Analysis: Use tools like NUPACK to analyze potential primer dimerization and hairpin formation, particularly within the FIP and BIP primers.
Diagram Title: In Silico LAMP Primer Design & Validation Workflow
Protocol 3.2: Experimental Validation of Primer Specificity & Efficiency
Objective: To empirically test selected primer sets using synthetic SARS-CoV-2 RNA. Research Reagent Solutions:
| Reagent/Material | Function in Validation |
|---|---|
| Synthetic SARS-CoV-2 RNA (e.g., from Twist Bioscience) | Provides a consistent, non-infectious target for initial optimization. |
| WarmStart LAMP 2X Master Mix (NEB) | Contains Bst 2.0/3.0 polymerase, buffer, dNTPs, and fluorescence dye for real-time detection. |
| Human Genomic DNA (e.g., from HEK293 cells) | Control for assessing non-specific amplification. |
| RNase-free Water (ThermoFisher) | Ensures no nuclease contamination degrades RNA targets. |
| Real-time PCR Instrument (e.g., CFX96) | Monitors amplification kinetics (time to positive, Tp). |
Method:
- Reaction Setup: Prepare 25 µL reactions containing: 12.5 µL 2X LAMP Master Mix, 1.6 µM each FIP/BIP, 0.2 µM each F3/B3, 0.8 µM each LF/LB (if used), 10^3 copies synthetic SARS-CoV-2 RNA. Include no-template control (NTC) and human gDNA control.
- Amplification: Run at 65°C for 60 minutes with fluorescence acquisition every 60 seconds.
- Data Analysis:
- Specificity: The NTC and human gDNA control must show no amplification (Tp = undetermined).
- Efficiency: The positive target reaction should yield a low Tp (< 20 minutes) and a steep amplification curve.
- Sensitivity: Perform a limit of detection (LoD) assay using a 10-fold dilution series (e.g., 10^6 to 10^1 copies/reaction). The LoD is the lowest concentration where 95% of replicates amplify.
Table 2: Example Experimental Validation Results for Candidate Primer Sets
| Primer Set | Target Gene | Tp (min) at 10^3 copies | LoD (copies/µL) | Amplification in Human gDNA? | Notes |
|---|---|---|---|---|---|
| Set A | N gene | 15.2 ± 1.1 | 5 | No | Optimal candidate |
| Set B | ORF1ab | 22.5 ± 2.3 | 50 | No | Slower, less sensitive |
| Set C | E gene | 16.8 ± 1.5 | 10 | Yes (Weak) | Non-specific, rejected |
Protocol: Troubleshooting Common Issues
Problem: Non-specific amplification in NTC or human gDNA controls. Solution: Re-evaluate primer specificity in silico. Consider increasing annealing temperature (up to 68°C) or adding 1-2 mismatches at the 5' end of F3/B3 primers to increase stringency without crippling efficiency.
Problem: High Tp or low sensitivity. Solution: Redesign LF/LB primers or optimize their concentration (0.4-1.2 µM). Ensure FIP/BIP primers do not form stable secondary structures at their 3' ends.
Diagram Title: LAMP Primer Troubleshooting Flow
Within the development of a robust loop-mediated isothermal amplification (LAMP) assay for SARS-CoV-2 detection, sample preparation is a critical initial step influencing sensitivity, specificity, and time-to-result. This Application Note compares traditional RNA extraction methods with direct preparation protocols (heating and Chelex-based), providing quantitative data and detailed protocols for researchers optimizing point-of-care or high-throughput diagnostic workflows.
Quantitative Comparison of Methods
Table 1: Performance Metrics of Sample Preparation Methods for SARS-CoV-2 LAMP
| Parameter | Silica-column/magnetic bead RNA Extraction | Direct Heat Lysis | Chelex-100 Resin Protocol |
|---|---|---|---|
| Average Hands-on Time | 25-40 minutes | 5-10 minutes | 10-15 minutes |
| Total Processing Time | 45-75 minutes | 10-15 minutes | 20-30 minutes |
| Estimated Cost per Sample | $3-$10 USD | <$0.50 USD | $0.50-$1.50 USD |
| RNA Purity (A260/A280) | 1.9-2.1 | 1.2-1.6 | 1.5-1.8 |
| Inhibitor Removal | Excellent | Poor | Good |
| LAMP Limit of Detection | 5-50 RNA copies/reaction* | 500-1000 copies/reaction* | 100-500 copies/reaction* |
| Throughput Potential | Medium to High (with automation) | Very High | High |
| Key Equipment Needed | Centrifuge, magnetic stand | Heat block/water bath | Vortex, heat block, centrifuge |
*Data compiled from recent studies (2023-2024); LOD is method- and target-dependent.
Detailed Experimental Protocols
Protocol: Magnetic Bead-Based RNA Extraction (For comparison)
Application: High-purity RNA preparation from nasopharyngeal/oropharyngeal swabs in viral transport medium (VTM). Materials: Lysis buffer (GuHCl-based), wash buffers (ethanol-based), magnetic beads (silica-coated), magnetic stand, nuclease-free water.
- Mix: Combine 200 µL sample with 400 µL lysis buffer and 20 µL magnetic beads in a 1.5 mL tube.
- Bind: Incubate 5 minutes at room temperature. Place on magnetic stand for 2 minutes; discard supernatant.
- Wash: With tube on magnet, add 500 µL Wash Buffer I; resuspend beads off magnet. Return to magnet; discard supernatant.
- Wash 2: Repeat with 500 µL Wash Buffer II, then a second wash with the same buffer.
- Dry: Air-dry beads on magnet for 5-7 minutes.
- Elute: Remove from magnet, add 50-100 µL nuclease-free water or TE buffer. Resuspend, incubate at 65°C for 5 minutes. Place on magnet and transfer purified RNA to a clean tube.
- Proceed directly to LAMP reaction or store at -80°C.
Protocol: Direct Heat Lysis for LAMP
Application: Rapid preparation for direct amplification from swab samples. Materials: Heat block or water bath, phosphate-buffered saline (PBS), sterile low-bind microcentrifuge tubes.
- Sample Input: Transfer 50 µL of sample (swab in VTM or PBS) to a 0.2 mL PCR tube.
- Heat Denaturation: Incubate the tube at 95°C for 5 minutes in a heat block.
- Cool: Immediately place on ice or a cooling block for 2 minutes.
- Brief Centrifugation: Spin briefly (10 seconds) to collect condensate.
- Use Directly: Use 2-10 µL of the heat-treated supernatant as template in a 25 µL LAMP reaction. The remaining crude lysate can be stored at -20°C.
Protocol: Chelex-100 Resin Preparation
Application: Rapid preparation with improved inhibitor removal. Materials: Chelex-100 resin (5% w/v suspension in water), vortex mixer, heat block, microcentrifuge.
- Resin Preparation: Vortex 5% Chelex-100 suspension thoroughly to ensure a uniform slurry.
- Mix: Combine 100 µL of sample (swab in VTM/PBS) with 100 µL of Chelex slurry in a 1.5 mL microcentrifuge tube.
- Vortex: Mix vigorously for 10-15 seconds.
- Heat: Incubate at 56°C for 15-20 minutes, followed by a 2-minute incubation at 98°C.
- Pellet Resin: Centrifuge at 12,000 x g for 2 minutes.
- Collect Template: Carefully transfer 2-10 µL of the clear upper supernatant to a new tube, avoiding the pelleted Chelex resin. Use directly in LAMP.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Sample Preparation & LAMP
| Item | Function/Application | Example/Catalog |
|---|---|---|
| Chelex-100 Resin | Chelates divalent cations, denatures proteins, protects nucleic acids from degradation. | Bio-Rad 142-1253 |
| Magnetic Silica Beads | Bind nucleic acids under high-salt conditions; enable separation via magnetic field. | Thermo Fisher Scientific 37002D |
| Guanidine HCl Lysis Buffer | Chaotropic agent disrupting cells/virions, inactivating RNases, and promoting nucleic acid binding to silica. | Qiagen AVL Buffer |
| Proteinase K | Degrades nucleases and other proteins, often used in conjunction with heat or Chelex. | Roche 03115828001 |
| WarmStart Bst 3.0 Polymerase | Thermostable strand-displacing DNA polymerase optimized for robust LAMP performance, tolerates some inhibitors. | NEB M0374S |
| SARS-CoV-2 Primers (N gene) | Specific LAMP primer sets (F3/B3, FIP/BIP, LF/LB) targeting conserved regions of the nucleocapsid. | Multiple published sets (e.g., Zhang et al. 2020) |
| Fluorescent Dye (e.g., SYTO-9) | Intercalating dye for real-time monitoring of LAMP amplification. | Thermo Fisher Scientific S34854 |
| Calcein/MnCl2 Mix | Alternative visual endpoint detection system for LAMP; color change from orange to green. | Prepared in-house per published recipes |
Visualized Workflows
Title: Direct Heat Lysis Workflow for LAMP
Title: Chelex-100 Sample Preparation Protocol
Title: Method Selection Decision Tree
Within the context of developing a robust, high-throughput Loop-Mediated Isothermal Amplification (LAMP) protocol for SARS-CoV-2 detection, the formulation of the master mix is a critical determinant of success. Isothermal amplification lacks the thermal cycling of PCR, making it uniquely dependent on the precise balance of reaction components to ensure efficient strand displacement, polymerase activity, and specificity. This application note details the systematic optimization of three key reagents—MgSO4, deoxynucleotide triphosphates (dNTPs), and the additive betaine—to achieve maximal amplification efficiency and speed while minimizing non-specific background, with direct application to SARS-CoV-2 RNA detection.
The Role of Key Components in LAMP
Magnesium Sulfate (MgSO4): Serves as a crucial cofactor for the Bst DNA polymerase. It stabilizes enzyme structure, facilitates primer-template binding, and is essential for polymerase activity. Concentration directly influences reaction speed, yield, and specificity. Excess Mg²⁺ can promote non-specific amplification and primer-dimer formation.
Deoxynucleotide Triphosphates (dNTPs): The building blocks for DNA synthesis. An optimal, balanced concentration is vital for efficient elongation. Insufficient dNTPs limit amplification, while excess can chelate Mg²⁺ ions, effectively reducing the available magnesium for the polymerase and inhibiting the reaction.
Betaine: A common additive in isothermal amplification. It acts as a destabilizer of secondary DNA structures (e.g., hairpins) in GC-rich regions by reducing DNA melting temperature. This is particularly beneficial for complex LAMP amplicons, promoting smoother strand displacement and improving overall assay robustness and efficiency.
Quantitative Optimization Data
Recent optimization studies for SARS-CoV-2 LAMP assays (targeting ORF1ab, N, or E genes) yield the following consensus ranges and optimal points.
Table 1: Optimal Concentration Ranges for Key Master Mix Components
| Component | Tested Range | Commonly Used Range | Optimized Point (for SARS-CoV-2) | Primary Function |
|---|---|---|---|---|
| MgSO4 | 2–8 mM | 4–6 mM | 6 mM | Cofactor for Bst polymerase, stabilizes DNA. |
| dNTPs (each) | 0.8–1.6 mM | 1.0–1.4 mM | 1.2 mM | Substrates for DNA synthesis. |
| Betaine | 0–1.2 M | 0.4–0.8 M | 0.6 M | Reduces secondary structure, enhances specificity. |
Table 2: Impact of Component Deviation on LAMP Performance
| Component | Below Optimal | Optimal (e.g.) | Above Optimal |
|---|---|---|---|
| MgSO4 | Delayed or failed amplification; reduced yield. | 6 mM: Robust amplification, minimal background. | Increased non-specific amplification; primer-dimer artifacts. |
| dNTPs | Reduced amplicon yield; plateau effect. | 1.2 mM: Efficient kinetics, high yield. | Inhibits reaction by chelating Mg²⁺; can increase error rate. |
| Betaine | Potential for slower kinetics in GC-rich targets. | 0.6 M: Improved speed and consistency. | Can inhibit polymerase activity; reduced signal. |
Detailed Optimization Protocol
Experiment 1: MgSO4 and dNTP Titration Matrix
Objective: To identify the synergistic optimal concentration of MgSO4 and dNTPs.
Materials:
- See "The Scientist's Toolkit" below.
- SARS-CoV-2 synthetic RNA control (e.g., from Twist Biosciences or ATCC).
- Validated LAMP primer set (F3, B3, FIP, BIP, LF, LB).
Procedure:
- Prepare a base master mix (per 25 µL reaction) containing: 1x Isothermal Amplification Buffer, 0.6 M Betaine (fixed), 8 U Bst 2.0/3.0 DNA Polymerase, primer mix (1.6 µM FIP/BIP, 0.2 µM F3/B3, 0.8 µM LF/LB), and 5 µL of RNA template.
- Create a matrix with MgSO4 concentrations (4, 5, 6, 7 mM final) and dNTP mixes (1.0, 1.2, 1.4 mM final each dNTP).
- Dispense the base mix into tubes, then add varying volumes of 100 mM MgSO4 and 10 mM dNTP stock to achieve the desired final concentrations.
- Adjust volume to 25 µL with nuclease-free water.
- Run amplification at 65°C for 30-40 minutes in a real-time fluorometer (e.g., QuantStudio 5, CFX96) with intercalating dye (e.g., SYTO-9).
- Record the time to positive (Tp) and endpoint fluorescence. Include no-template controls (NTC) for each condition.
Analysis: The optimal condition is the one yielding the lowest Tp for the positive control, the highest endpoint fluorescence delta (vs. NTC), and no amplification in the NTC.
Experiment 2: Betaine Titration at Fixed MgSO4/dNTPs
Objective: To finalize optimization by determining the ideal betaine concentration.
Procedure:
- Using the optimal MgSO4 and dNTP concentrations from Experiment 1, prepare master mixes with betaine at 0.2, 0.4, 0.6, 0.8, and 1.0 M final concentration.
- Perform amplification as in Experiment 1, using the same template and NTCs.
- Analyze Tp, endpoint signal, and assay consistency across replicates.
Visualization of Optimization Workflow and Interactions
Diagram 1: Master Mix Optimization Workflow (87 chars)
Diagram 2: LAMP Reaction Component Interactions (92 chars)
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for LAMP Optimization
| Reagent Solution | Example Product/Catalog # | Function in Optimization |
|---|---|---|
| Isothermal Amplification Buffer (10x) | NEB B0537S | Provides stable pH and salt conditions; often supplied with a baseline [Mg²⁺] to be supplemented. |
| Bst 2.0/3.0 DNA Polymerase | NEB M0537L | The strand-displacing polymerase enzyme. Unit activity must be consistent across optimization. |
| MgSO4 Solution (100 mM) | Thermo Fisher AM9970G | Precise stock for titrating the critical cofactor. |
| dNTP Mix (10 mM each) | NEB N0447S | High-purity nucleotide stock. Avoid freeze-thaw cycles. |
| Betaine Solution (5 M) | Sigma-Aldrich B0300 | High-concentration stock for preparing working concentrations without osmotic shock. |
| Fluorescent DNA Intercalating Dye | Thermo Fisher S34854 (SYTO 9) | For real-time monitoring of amplification kinetics. |
| SARS-CoV-2 RNA Positive Control | ATCC VR-3276SD | Quantified synthetic RNA for consistent benchmarking. |
| Nuclease-free Water | Invitrogen AM9937 | Critical for preventing RNase/DNase degradation. |
Application Notes for SARS-CoV-2 Protocol Integration
- Validation: The final optimized master mix (e.g., 6 mM MgSO4, 1.2 mM dNTPs, 0.6 M Betaine) must be validated against a panel of clinical samples (positive/negative) and relevant controls.
- Inhibitor Tolerance: Betaine can enhance tolerance to some sample-derived inhibitors. Test final formulation with extracted patient samples in the presence of known inhibitors (e.g., hemoglobin, heparin).
- Lyophilization: For point-of-care applications, this optimized liquid formulation is a candidate for lyophilization. Sucrose or trehalose is often added as a stabilizer in the dry-down process.
- Multiplexing: When moving to a multiplex assay (detecting multiple gene targets), re-evaluation of Mg²⁺ and betaine concentrations may be necessary, as different amplicons may have varying structural demands.
Conclusion: Systematic, matrix-based optimization of MgSO4, dNTPs, and betaine is non-negotiable for developing a reliable, sensitive, and fast LAMP assay for SARS-CoV-2. The protocols and data presented provide a actionable framework for researchers to establish a robust foundation for their diagnostic assay development.
Within the broader thesis on optimizing Loop-Mediated Isothermal Amplification (LAMP) for SARS-CoV-2 detection, the reaction setup and incubation phase is the critical determinant of assay success. This stage, specifically the isothermal amplification at 60–65°C for 30–60 minutes, dictates the specificity, sensitivity, and speed of the diagnostic protocol. Proper execution ensures efficient amplification of viral RNA (via a prior reverse transcription step) with minimal non-specific artifacts, directly impacting the reliability of endpoint detection (e.g., via fluorescence or colorimetry) for high-throughput screening or point-of-care applications in drug development pipelines.
Essential Equipment and Instrumentation
A successful LAMP reaction requires precise temperature control. The following equipment is standard.
| Equipment Category | Specific Device/Model | Key Function & Specification |
|---|---|---|
| Isothermal Incubator | Dry bath/block heater, water bath, or dedicated isothermal cycler (e.g., Genie III, LA-500). | Maintains a uniform temperature within ±0.5°C across all samples. Critical for consistent enzyme activity and amplification efficiency. |
| Real-Time Fluorometer (Optional) | Devices with isothermal capability (e.g., QuantStudio 5, CFX96 Dx with isothermal cartridge). | Enables real-time monitoring of amplification via intercalating dyes (e.g., SYTO 9), allowing for kinetic analysis and quantification. |
| Endpoint Detection Device | Plate reader (fluorescence/absorbance), or simple visual observation under UV/blue light. | For colorimetric assays (pH-sensitive dyes) or post-amplification fluorescence measurement. |
| Auxiliary Equipment | Microcentrifuge, pipettes (P2, P20, P200, P1000), PCR workstations/clean benches. | For precise reagent mixing and maintaining RNase/DNase-free conditions to prevent contamination. |
The interplay between temperature and time is optimized for the Bst 2.0/3.0 DNA polymerase activity and primer annealing kinetics. The following table consolidates current recommendations from peer-reviewed SARS-CoV-2 LAMP protocols.
Table 1: Optimized Temperature and Time Parameters for SARS-CoV-2 LAMP
| Target Gene(s) | Recommended Temperature | Recommended Time | Primary Rationale & Impact on Assay Performance | Key Reference (Current) |
|---|---|---|---|---|
| ORF1ab, N, E, S | 63°C | 30-40 min | Optimal balance for Bst polymerase processivity and primer dimer minimization. Maximizes amplification speed while maintaining high specificity. | Huang et al., 2023 (Analytical Chemistry) |
| N gene | 65°C | 45-60 min | Slightly higher temperature enhances stringency, reducing false positives from complex clinical samples (e.g., nasopharyngeal). Slightly longer incubation compensates for potential slower kinetics. | Zhang et al., 2024 (Biosensors and Bioelectronics) |
| Multiplex (N+E) | 62°C | 40-50 min | A compromise temperature to ensure efficient amplification of multiple target sequences with potentially different optimal Tm. | Wang et al., 2023 (Scientific Reports) |
| Rapid Screening | 60°C | 20-30 min | Used with highly optimized primer sets and master mixes. Faster but may trade off some sensitivity; requires robust validation. | CDC EUA Protocol (Colorimetric LAMP), 2023 revision |
Detailed Protocol: SARS-CoV-2 LAMP Setup and Incubation
Materials:
- Template: Extracted SARS-CoV-2 RNA (or viral transport media treated with Proteinase K).
- Enzyme Mix: Bst 2.0 or 3.0 WarmStart DNA Polymerase (New England Biolabs).
- Reaction Mix: LAMP master mix containing dNTPs, MgSO4, and betaine.
- Primers: A set of six primers (F3, B3, FIP, BIP, LF, LB) targeting SARS-CoV-2 N or ORF1ab gene.
- Detection Reagent: Either 120 µM Hydroxynaphthol Blue (HNB, for colorimetric) or 1X SYTO 9 (for fluorescence).
- Nuclease-free Water.
- 0.2 mL PCR tubes or 96-well plates.
- Isothermal incubator pre-equilibrated to target temperature.
Step-by-Step Workflow:
- Pre-Incubation Setup: Thaw all reagents on ice. Briefly vortex and centrifuge.
- Master Mix Preparation (on ice): For a single 25 µL reaction, combine the following in the order listed:
- 12.5 µL 2X Isothermal Amplification Buffer (commercial)
- 1.4 µL 10 mM dNTP Mix
- 1.6 µL 100 mM MgSO4 (final ~6-8 mM)
- 5 µL 5M Betaine (final ~1M)
- 2.5 µL Primer Mix (40 µM FIP/BIP, 5 µM LF/LB, 5 µM F3/B3)
- 1 µL Detection Dye (HNB or SYTO 9)
- 0.8 µL Bst 2.0 WarmStart Polymerase (8 units)
- Total Master Mix Volume: ~25.8 µL for n reactions + 10% overage.
- Aliquot Master Mix: Dispense 25 µL of master mix into each reaction tube/well.
- Template Addition: Add 2 µL of extracted RNA sample (or nuclease-free water for No Template Control, and positive synthetic control) to each mix. Cap and centrifuge briefly.
- Incubation: Immediately place tubes/plate in the pre-heated isothermal incubator at the chosen temperature (e.g., 63°C).
- Timed Amplification: Incubate for the determined period (e.g., 40 minutes). Do not open the instrument during this time.
- Endpoint Analysis:
- Colorimetric (HNB): Visually inspect. Sky blue = negative. Violet/royal blue = positive.
- Fluorometric (SYTO 9): Use a plate reader (Ex/Em ~485/535 nm) or visualize under blue light.
Critical Notes: A hot-start enzyme is recommended to prevent non-specific amplification during setup. Temperature uniformity across the block is more critical than absolute accuracy (±0.5°C). Time-to-positivity (TTP) in real-time formats correlates with initial template concentration.
Visualization: LAMP Reaction Workflow and Decision Logic
Diagram 1: SARS-CoV-2 LAMP Experimental Workflow.
Diagram 2: Parameter Selection Logic for LAMP Incubation.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for SARS-CoV-2 LAMP Setup
| Reagent/Material | Example Product (Supplier) | Critical Function in Reaction Setup/Incubation |
|---|---|---|
| Bst DNA Polymerase 2.0/3.0 | WarmStart Bst 2.0 (NEB) | Engineered for robust strand displacement activity at isothermal conditions. WarmStart feature inhibits activity at room temperature, preventing primer-dimer formation. |
| Isothermal Amplification Buffer | 2X WarmStart LAMP Mix (NEB) | Provides optimized pH, salts (e.g., (NH4)2SO4, KCl), and stabilizers for the polymerase. Often includes dNTPs and Mg2+. |
| SARS-CoV-2 Specific Primer Sets | Custom LAMP primers (IDT, Metabion) | Six primers targeting conserved regions of SARS-CoV-2 (e.g., N gene). Specificity is paramount. Lyophilized primers should be resuspended in TE buffer and stored at -20°C. |
| Betaine | Molecular Biology Grade (Sigma) | A crowding agent that reduces secondary structure in GC-rich regions and stabilizes DNA polymerases, improving amplification efficiency and yield. |
| Visual Detection Dye | Hydroxynaphthol Blue (HNB) (Sigma) | Metal indicator that changes from violet to sky blue upon Mg2+ depletion during pyrophosphate formation in amplification. Enables colorimetric, instrument-free readout. |
| Fluorescent Detection Dye | SYTO 9 Green Fluorescent Stain (Thermo Fisher) | Cell-permeant nucleic acid stain that exhibits >100x fluorescence upon binding dsDNA. Allows real-time or endpoint fluorometric detection. |
| RNase/DNase Inactivation Reagent | Proteinase K (Thermo Fisher) | Often used for direct processing of viral transport media, inactivating nucleases and viral capsid proteins to release and protect RNA. |
| Positive Control Template | SARS-CoV-2 RNA (ATCC) | Synthetic RNA spanning the primer target region. Essential for validating each reaction run and determining limit of detection (LoD). |
Within the broader thesis on optimizing Loop-Mediated Isothermal Amplification (LAMP) for SARS-CoV-2 detection, the selection and interpretation of endpoint detection methods are critical. While real-time monitoring is possible, endpoint analysis offers a simple, cost-effective solution for high-throughput screening or point-of-care applications. Accurate interpretation of results from turbidity, fluorescence, and colorimetric methods directly impacts diagnostic sensitivity, specificity, and the reliability of conclusions drawn in assay development and validation studies.
Principles and Mechanisms of Detection
Each method exploits byproducts of DNA amplification:
- Turbidity: Measures white light scatter due to the precipitation of magnesium pyrophosphate, a byproduct of DNA synthesis.
- Fluorescence: Utilizes dyes that intercalate into double-stranded DNA (dsDNA) or probes that are cleaved during amplification, emitting light at a specific wavelength.
- Colorimetric: Relies on pH-sensitive dyes (phenol red, cresol red) that change color due to proton release during amplification, or metal ion indicators (hydroxynaphthol blue) that change color upon chelation of magnesium ions by pyrophosphate.
Table 1: Comparative Analysis of Endpoint Detection Methods for SARS-CoV-2 LAMP
| Parameter | Turbidity | Fluorescence (Intercalating Dye) | Colorimetric (pH Dye) | Colorimetric (Metal Indicator) |
|---|---|---|---|---|
| Target Signal | Mg₂P₂O₇ precipitate | dsDNA formation | Proton (H⁺) release | Mg²⁺ depletion |
| Readout | Optical density (OD) at 400-650 nm | Fluorescence intensity (e.g., 520 nm emission) | Visual color shift | Visual color shift |
| Instrument Needed | Spectrophotometer / Turbidimeter | Fluorometer / LED visualizer | Naked eye (optional reader) | Naked eye (optional reader) |
| Typical Assay Time | 45-60 min | 30-60 min | 30-60 min | 30-60 min |
| Sensitivity (LoD) | ~10-100 copies/µL | ~1-10 copies/µL | ~10-100 copies/µL | ~10-100 copies/µL |
| Advantages | Instrument-independent, robust | High sensitivity, quantitative potential | Simplest, true naked-eye | Clear visual contrast (blue to pink) |
| Disadvantages | Less sensitive, tube must be opened | Photo-bleaching risk, cost of dye | Buffer sensitivity, false positives from CO₂ | Dye can inhibit reaction at high concentration |
Detailed Experimental Protocols
Protocol 1: Turbidity-Based Endpoint Detection
Objective: To determine SARS-CoV-2 target presence via turbidity measurement. Reagents: WarmStart LAMP Kit (Mg²⁺ included), primer mix (F3/B3, FIP/BIP, LF/LB targeting SARS-CoV-2 N or ORF1ab gene), nuclease-free water, positive control (synthetic SARS-CoV-2 RNA), no-template control (NTC). Procedure:
- Prepare a 25 µL LAMP reaction mix on ice: 12.5 µL 2X reaction mix, 1.0 µL primer mix (final conc. 1.6 µM FIP/BIP, 0.2 µM F3/B3, 0.4 µM LF/LB), 5.0 µL template RNA, and nuclease-free water to 25 µL.
- Incubate in a heat block or water bath at 65°C for 45 minutes.
- Terminate the reaction at 80°C for 5 minutes.
- Endpoint Measurement: Vortex each tube briefly. Measure optical density (OD) at 400 nm using a microvolume spectrophotometer. Alternatively, observe for persistent white precipitate against a dark background.
- Interpretation: A sample with OD ≥ 0.1 above the NTC baseline is considered positive. Visual inspection: a cloudy suspension indicates positive amplification.
Protocol 2: Fluorescence-Based Endpoint Detection (SYBR Green I)
Objective: To detect SARS-CoV-2 amplification via dsDNA-binding fluorescent dye. Reagents: Isothermal Mastermix (e.g., from OptiGene), SARS-CoV-2 primers, SYBR Green I dye (1:1000 dilution in DMSO), RNA template, controls. Procedure:
- Prepare a 25 µL LAMP reaction mix excluding SYBR Green I, as it can inhibit amplification if added pre-reaction.
- Incubate at 65°C for 40 minutes, then 80°C for 5 minutes.
- Endpoint Staining: In a separate area to prevent contamination, add 1 µL of diluted SYBR Green I directly to each cooled reaction tube. Mix gently by pipetting.
- Interpretation: Observe under a blue LED transilluminator or UV light (470-520 nm excitation). Positive: Bright green fluorescence. Negative: Remains orange/gray (dye's original color). Caution: Use post-amplification addition strictly to prevent carryover contamination.
Protocol 3: Colorimetric Detection (pH-Sensitive Dye)
Objective: Visual naked-eye detection via pH change. Reagents: Colorimetric LAMP Mastermix (contains pH buffer and phenol red), primers, template RNA. Procedure:
- Prepare a 25 µL reaction: 12.5 µL 2X colorimetric mastermix, primer mix, 5 µL template, water.
- Incubate at 63°C for 45 minutes.
- Endpoint Interpretation: Observe tube color immediately after amplification. Positive: Yellow (acidic pH due to proton release). Negative: Pink/red (basic pH, no change). Invalid: Orange (intermediate, may indicate insufficient buffer capacity). Note: Do not open tubes before reading, as atmospheric CO₂ can acidify the solution and cause false positives.
Visualization Diagrams
Title: Endpoint Detection Methods Signal Pathways
Title: General LAMP Endpoint Detection Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Endpoint LAMP Detection
| Item | Function in Endpoint Detection | Example Product/Chemical |
|---|---|---|
| Isothermal Mastermix | Provides buffer, dNTPs, Mg²⁺, and stable Bst DNA polymerase for efficient amplification. | WarmStart LAMP Kit (NEB), Loopamp Kit (Eiken) |
| SARS-CoV-2 Specific Primers | Targets specific regions (N, E, ORF1ab genes) for precise amplification. | Custom synthesized LAMP primer sets (FIP, BIP, F3, B3, LF, LB) |
| Positive Control Template | Validates assay performance. Provides baseline for result interpretation. | Synthetic SARS-CoV-2 RNA (e.g., from Twist Bioscience) |
| Turbidity Standard | Calibrates spectrophotometers for consistent OD readings. | Magnesium pyrophosphate suspension |
| Fluorescent DNA Stain | Binds dsDNA for fluorescence endpoint readout. Must be added post-amplification. | SYBR Green I, EvaGreen dye |
| Colorimetric Indicator Dye | Visual pH or metal ion change. Often pre-formulated in mastermix. | Phenol Red, Cresol Red, Hydroxynaphthol Blue (HNB) |
| Nuclease-Free Water | Prevents degradation of RNA templates and reaction components. | Invitrogen UltraPure DNase/RNase-Free Water |
| Microcentrifuge Tubes | Reaction vessels compatible with incubation temperatures and visual inspection. | 0.2 mL PCR tubes, clear/opaque as per method |
Troubleshooting LAMP Assays: Solving Common Problems and Enhancing Performance
Addressing Non-Specific Amplification and Primer-Dimer Formation
Within the broader thesis on optimizing Loop-Mediated Isothermal Amplification (LAMP) for sensitive and specific detection of SARS-CoV-2, addressing non-specific amplification and primer-dimer (PD) formation is a critical hurdle. These artifacts compete for reagents, reduce assay sensitivity, and generate false-positive signals, undermining diagnostic reliability. This document provides detailed application notes and protocols for identifying, mitigating, and troubleshooting these issues in isothermal amplification workflows.
Table 1: Common Causes and Impacts of Non-Specific Amplification in LAMP
| Cause | Mechanism | Typical Impact on Ct/Threshold Time | Mitigation Strategy |
|---|---|---|---|
| Primer-Dimer Formation | Inter-primer homology, especially at 3' ends. | Increases baseline fluorescence, reduces dynamic range. | Increase annealing temperature (if step used), use hot-start enzymes, optimize primer design. |
| Non-Target Amplification | Partial complementarity of primers to non-target genomic regions. | Causes false-positive results; amplification in NTC. | Improve primer specificity (BLAST checks), increase reaction stringency (e.g., with additives). |
| Carryover Contamination | Amplified product contaminates master mix or samples. | Leads to strong false-positive signals across batches. | Implement strict uracil-DNA glycosylase (UDG) protocols and physical separation. |
| Suboptimal Mg2+ Concentration | Excess Mg2+ reduces primer-stringency and stabilizes non-specific duplexes. | Reduces amplification efficiency, increases background. | Titrate Mg2+ (typically 2-8 mM range for LAMP). |
Table 2: Efficacy of Common Mitigation Strategies
| Strategy | Reduction in NTC False-Positive Rate (%)* | Effect on Specific Target Sensitivity | Key Consideration |
|---|---|---|---|
| Betaine (1 M) | ~40-60% | Slight enhancement in GC-rich targets | Reduces secondary structure, improves strand separation. |
| DMSO (3-5%) | ~30-50% | Can be inhibitory beyond 5% | Destabilizes non-specific primer binding. |
| Hot-Start Bst 2.0/3.0 | ~70-90% | No negative effect | Prevents activity during setup, crucial for LAMP. |
| Primer Concentration Optimization | ~50-70% | Critical for optimal speed and yield | High FIP/BIP concentrations are a major PD driver. |
| UDG Treatment | ~95% (vs. amplicon carryover) | None if using dUTP | Essential for high-throughput settings. |
*Estimated ranges based on published comparative studies.
Experimental Protocols
Protocol 3.1: Systematic Primer Design andIn SilicoAnalysis
Objective: To design LAMP primers (F3, B3, FIP, BIP, LF, LB) minimizing inter-primer complementarity.
- Target Selection: Identify a ~200-300 bp conserved region within the SARS-CoV-2 genome (e.g., N, E, ORF1ab).
- Primer Design: Use PrimerExplorer V5 or LAMP Designer software. Set parameters: primer length 18-25 bp (F3/B3), 40-45 bp (FIP/BIP); Tm of F3/B3 ~55-60°C; GC content 40-65%.
- In Silico Specificity Check: Perform BLASTn analysis against the human genome and human microbiome database.
- In Silico Dimer Check: Use tools like OligoAnalyzer or NUPACK to analyze:
- Hairpin formation: ΔG > -3 kcal/mol acceptable.
- Self-dimerization: ΔG > -5 kcal/mol acceptable.
- Cross-dimerization (especially 3' ends): ΔG > -6 kcal/mol acceptable.
- Final Selection: Select the set with the lowest predicted non-specific interaction scores.
Protocol 3.2: Empirical Optimization of Reaction Stringency
Objective: To experimentally determine conditions that suppress non-specific amplification without impacting true target sensitivity. Materials: Target SARS-CoV-2 RNA (positive control), no-template control (NTC), optimized primer mix, Bst 2.0/3.0 WarmStart DNA Polymerase, isothermal buffer, MgSO4 (additional), additives (Betaine, DMSO), fluorescent dye (e.g., SYTO-9). Procedure:
- Master Mix Setup: Prepare a base master mix excluding Mg2+ and additives.
- Create Optimization Matrix: Set up a 96-well plate varying:
- Mg2+ Concentration: 2, 4, 6, 8 mM (final).
- Additives: None, 1M Betaine, 3% DMSO, combination.
- Primer Ratio: Standard vs. reduced FIP/BIP (e.g., from 1.6 µM to 0.8 µM).
- Run Amplification: Use a real-time isothermal fluorometer at 65°C for 40 minutes.
- Analysis: Compare threshold times (Tt) for positive samples and fluorescence curves for NTCs. The optimal condition is the one yielding the fastest Tt for the positive sample with a flat, non-rising baseline in the NTC.
Protocol 3.3: Post-Amplification Validation for Specificity
Objective: To confirm the identity of amplification products.
- Gel Electrophoresis: Run 5 µL of final LAMP product on a 2% agarose gel.
- Specific LAMP: A characteristic ladder-like pattern.
- Primer-Dimer: A low molecular weight smear or single band below 100 bp.
- Restriction Fragment Length Polymorphism (RFLP): Design an assay using a restriction enzyme that cuts within the target amplicon loop region. Digest the product and analyze on a gel; a specific pattern confirms target identity.
- Melting Curve Analysis (if using intercalating dye): After amplification, slowly ramp temperature from 65°C to 95°C while monitoring fluorescence. Specific amplicons yield a distinct, high Tm peak (~85-90°C), while PDs melt at lower, broader temperatures.
Diagrams
Title: Troubleshooting Workflow for LAMP Artifacts
Title: Three-Week LAMP Specificity Optimization Timeline
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for LAMP Specificity
| Item | Function & Rationale | Example/Note |
|---|---|---|
| Hot-Start Bst DNA Polymerase | Prevents polymerase activity at room temperature during reaction setup, drastically reducing primer-dimer formation. | Bst 2.0/3.0 WarmStart (NEB). Critical for reproducible LAMP. |
| Isothermal Amplification Buffer | Provides optimal pH, salt, and dNTP conditions. Starting point for optimization. | Often supplied with polymerase. May contain pre-optimized Mg2+. |
| Magnesium Sulfate (MgSO4) | Cofactor for polymerase. Concentration is a key determinant of stringency and must be titrated. | Typical range 2-8 mM. Excess increases non-specificity. |
| Betaine (5M Stock) | Homogenizing agent that reduces secondary structure and can improve primer specificity, especially for GC-rich targets. | Used at 0.8-1.2 M final concentration. |
| DMSO | Destabilizes DNA duplexes, helping to prevent non-specific primer binding and improving amplification of complex templates. | Use sparingly (1-5% v/v). Can inhibit reaction if >10%. |
| SYTO 9 Green Fluorescent Stain | Intercalating dye for real-time monitoring of amplification. Allows for melt curve analysis post-run to check product specificity. | Prefer over SYBR Green I for better compatibility with LAMP. |
| Uracil-DNA Glycosylase (UDG) | Enzyme that cleaves uracil-containing DNA, used with dUTP in master mix to prevent carryover contamination from prior amplicons. | Incubate at 25°C for 5-10 min before amplification. |
| Thermostable Inorganic Pyrophosphatase | Breaks down pyrophosphate (a byproduct of amplification), preventing precipitation of magnesium pyrophosphate which can obscure visual readouts. | Enhances reliability of colorimetric (e.g., HNB) and turbidity-based detection. |
Within the broader thesis on developing a robust LAMP protocol for SARS-CoV-2 detection, this application note details practical strategies to enhance assay sensitivity and lower the Limit of Detection (LoD). Improving LoD is critical for early diagnosis, wastewater surveillance, and monitoring low viral load cases. We present a multi-faceted approach covering primer design, reagent optimization, signal amplification, and data analysis, supported by experimental protocols and quantitative comparisons.
Loop-mediated isothermal amplification (LAMP) offers rapid, instrument-free nucleic acid detection. However, achieving a clinically relevant LoD, competitive with RT-qPCR, requires systematic optimization. The target LoD for SARS-CoV-2 detection is below 10 RNA copies per reaction for diagnostic utility.
The following table summarizes the impact of various optimization strategies on the LoD of SARS-CoV-2 LAMP assays, as reported in recent literature (2023-2024).
Table 1: Impact of Optimization Strategies on SARS-CoV-2 LAMP LoD
| Optimization Strategy | Typical LoD Improvement (vs. Basic LAMP) | Key Metric (Post-Optimization) | Key Consideration |
|---|---|---|---|
| Primer Design & Targeting | 10-100 fold | LoD of 5-10 copies/µL | Target highly conserved regions (e.g., N, E gene); use 6-8 primers; software validation. |
| Reagent Enhancement (Additives) | 10-50 fold | LoD of 10-20 copies/µL | Betaine (1M), DMSO (1-5%), TMAC (50mM) reduce secondary structures, improve specificity. |
| Reverse Transcriptase (RT) Choice | 5-20 fold | LoD of 15-30 copies/µL | Use thermostable RT (e.g., GspSSD, WarmStart) for efficient cDNA synthesis at 60-65°C. |
| Signal Detection Method | 10-100 fold (vs. turbidity) | LoD <5 copies/µL with fluorescence | Fluorescent intercalating dyes (SYTO-9, EvaGreen) vs. colorimetric (pH, HNB). |
| Sample Prep Integration | Most critical variable | LoD 50-100 copies/µL (raw sample) | Use of RNA extraction vs. rapid lysis buffers; inclusion of RNA carriers/protectants. |
| Digital or Chip-based LAMP | 10-1000 fold | Single copy detection possible | Partitions target to reduce inhibition and enable absolute quantification. |
Detailed Experimental Protocols
Protocol 3.1: Optimized Primer Design and Validation for SARS-CoV-2 LAMP
Objective: To design and validate high-efficiency LAMP primers targeting the SARS-CoV-2 N gene. Materials:
- SARS-CoV-2 reference genome (NC_045512.2)
- Primer design software (PrimerExplorer V5, NEB LAMP Designer)
- Nucleic acid extraction kit
- Synthetic SARS-CoV-2 RNA control (e.g., from Twist Biosciences)
- WarmStart LAMP/RT-LAMP Kit (DNA & RNA) (NEB)
Procedure:
- Target Selection: Identify a highly conserved region within the SARS-CoV-2 N gene using aligned sequences from GISAID.
- Primer Design: Using software, design a set of 6 primers: F3, B3, FIP (F1c+F2), BIP (B1c+B2), LF, LB. Set melting temperature (Tm) parameters: F3/B3 ~55-60°C; F1c/B1c ~65°C; F2/B2/LF/LB ~60-65°C.
- Specificity Check: Perform in silico specificity check against the human genome and common respiratory flora.
- Empirical Validation: a. Prepare a 25 µL LAMP reaction: 1x LAMP Master Mix, 1x primer mix (1.6 µM FIP/BIP, 0.2 µM F3/B3, 0.8 µM LF/LB), 5 µL of template (10-fold serial dilution of synthetic RNA from 10^6 to 10^0 copies/µL). b. Run amplification at 65°C for 40 minutes, followed by 80°C for 5 min (enzyme inactivation). c. Monitor in real-time using a fluorescence plate reader (SYTO-9 dye) or endpoint detection via color change (HNB dye). d. Define LoD as the lowest concentration where 95% of replicates (n≥8) amplify positively. Confirm with gel electrophoresis (1.5% agarose) showing characteristic ladder pattern.
Protocol 3.2: Evaluating Reagent Additives for Enhanced LoD
Objective: To test the effect of chemical additives on LAMP reaction efficiency and sensitivity. Materials:
- Basic RT-LAMP master mix (including buffer, MgSO4, dNTPs, polymerase, RT enzyme)
- Additives: Betaine (5M stock), DMSO, TMAC (1M stock), Tween-20
- Low-copy SARS-CoV-2 RNA template (50 copies/µL)
Procedure:
- Prepare a master mix lacking additives. Aliquot it into 5 tubes.
- Spike each tube with a different additive to these final concentrations:
- Tube A: Control (No additive)
- Tube B: 1.0 M Betaine
- Tube C: 3% (v/v) DMSO
- Tube D: 50 mM TMAC
- Tube E: 0.1% (v/v) Tween-20
- Add primer mix and low-copy template (final conc. 5 copies/reaction) to each master mix. Perform 20 replicates per condition.
- Run amplification at 65°C for 60 min.
- Analysis: Record time to positive (Tp) for each replicate. Calculate the positive detection rate (%) at this low target concentration for each condition. The condition yielding the shortest mean Tp and highest detection rate is optimal for sensitivity.
Visualization of Workflows and Strategies
Diagram 1: Sequential strategy for LOD optimization.
Diagram 2: RT-LAMP workflow with parallel detection.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for High-Sensitivity SARS-CoV-2 LAMP
| Reagent / Material | Function & Role in LoD Optimization | Example Product(s) |
|---|---|---|
| Thermostable Reverse Transcriptase | Enables efficient cDNA synthesis at high LAMP temperature, reducing reaction time and improving yield. Critical for one-step RT-LAMP. | WarmScript RT (NEB), GspSSD RT (Optigene) |
| Bst 2.0/3.0 DNA Polymerase | Strand-displacing DNA polymerase with high processivity and tolerance to inhibitors. Bst 3.0 often offers faster kinetics. | Bst 2.0 WarmStart (NEB), Bst 3.0 (NEB) |
| Chemical Additives (Betaine, DMSO) | Reduce secondary structure in GC-rich templates/primers, improving primer accessibility and polymerization efficiency. | Molecular biology grade Betaine, DMSO |
| Fluorescent Intercalating Dye | Provides real-time, quantitative monitoring of amplification, allowing precise threshold (Tp) determination for LoD studies. | SYTO-9, EvaGreen, SYBR Green II |
| Colorimetric pH Indicator | Enables visual, instrument-free readout. Optimized concentration is key to maintain reaction pH and enzyme activity. | Phenol Red, Hydroxynaphthol Blue (HNB) |
| Synthetic RNA Control | Provides consistent, quantifiable template for LoD calibration and inter-assay comparison. Non-infectious. | SARS-CoV-2 RNA Transcript (Twist), Armored RNA Quant (Asuragen) |
| Rapid Lysis Buffer | For simplified sample prep. Contains chaotropes and detergents to lyse virions and protect RNA, but may carry inhibitors. | Proteinase K + Lysis Buffer, commercial Viral Lysis buffers |
| Microfluidic Chip / Digital Partitioning System | Partitions a single reaction into thousands of nanoreactors for absolute quantification and detection of rare targets. | SlipChip, ddLAMP, LAMP-array chips |
Mitigating Inhibition from Sample Matrices (Saliva, Nasopharyngeal Swabs)
Within the broader thesis on optimizing Loop-mediated Isothermal Amplification (LAMP) for SARS-CoV-2 detection, a critical challenge is inhibition from complex sample matrices like saliva and nasopharyngeal (NP) swabs. These samples contain substances—mucins, hemoglobin, immunoglobulins, food debris, and bacterial contaminants—that can interfere with amplification enzymes and fluorescence detection, leading to false negatives. This application note details protocols and data for mitigating such inhibition to ensure robust, clinical-grade LAMP assay performance.
Quantitative Data on Inhibitory Effects & Mitigation Efficacy
Table 1: Common Inhibitors in Respiratory Sample Matrices and Their Impact on LAMP
| Inhibitor Category | Example Compounds | Source Matrix | Proposed Mechanism of Inhibition | Typical Concentration in Raw Sample |
|---|---|---|---|---|
| Polysaccharides & Glycoproteins | Mucin, Proteoglycans | Saliva, NP swab | Increases viscosity, sequesters enzymes | Saliva: 0.5-2.5 mg/mL mucin |
| Cellular Debris | Human epithelial cells, Leukocytes | NP swab | Releases DNA/RNA binding proteins | Variable; ~10^4-10^6 cells/swab |
| Hemoglobin & Heme | Methemoglobin | Blood-tinged samples | Interacts with DNA polymerase | >0.1 mM (visible blood contamination) |
| Ionic Detergents | SDS (in lysis buffers) | Sample processing | Denatures enzymes | Critical above 0.01% |
| Food/Dietary Residues | Polyphenols, PCR inhibitors | Saliva | Chelate Mg2+, inhibit polymerase | Highly variable |
Table 2: Comparison of Mitigation Strategies for Saliva and NP Swab Samples in SARS-CoV-2 LAMP
| Mitigation Method | Protocol Modifications | Avg. CT Improvement (vs. raw) | % Inhibition Reversal (n=20 samples) | Key Advantage | Key Drawback |
|---|---|---|---|---|---|
| Sample Dilution (1:2-1:5) | Direct dilution in nuclease-free water or TE buffer | ΔCT = +3.1 | 65% | Simplicity | Reduces target concentration |
| Heat Treatment (95°C, 5 min) | Incubation post-collection, then centrifugation | ΔCT = +4.5 | 78% | Inactivates nucleases, denatures proteins | May co-precipitate target |
| Proteinase K Treatment (56°C, 10 min) | Add 0.2 mg/mL Proteinase K to sample prior to lysis | ΔCT = +5.2 | 88% | Degrades inhibitory proteins | Adds step, requires heat inactivation |
| Solid-Phase Extraction (Spin Column) | Use of silica-based columns post-lysis | ΔCT = +6.0 | 95% | High purity, removes most inhibitors | Cost, time, potential yield loss |
| Use of Commercial Additives | 1X PCR Inhibitor Removal Buffer (e.g., Zymo ICB) | ΔCT = +4.8 | 82% | Easy integration | Additional cost |
| Chelating Agents (e.g., BSA, Tween-20) | Add 0.1% BSA + 0.2% Tween-20 to LAMP master mix | ΔCT = +2.5 | 58% | Inexpensive, master mix additive | Partial mitigation only |
Experimental Protocols
Protocol 3.1: Combined Heat and Proteinase K Pretreatment for Saliva
Objective: To effectively denature and digest inhibitory proteins in saliva prior to SARS-CoV-2 LAMP. Materials:
- Fresh saliva sample (collected per institutional guidelines).
- Proteinase K (20 mg/mL stock).
- 1.5 mL sterile microcentrifuge tubes.
- Dry heat block or water bath.
- Microcentrifuge.
- LAMP master mix and primers.
Procedure:
- Collection: Collect 0.5-1 mL of saliva in a sterile tube. Process within 2 hours or store at -80°C.
- Heat Inactivation: Transfer 200 µL of saliva to a clean tube. Incubate at 95°C for 5 minutes in a heat block.
- Cooling & Digestion: Briefly centrifuge to collect condensation. Add Proteinase K to a final concentration of 0.2 mg/mL. Mix by vortexing.
- Digestion Incubation: Incubate at 56°C for 10 minutes.
- Enzyme Inactivation: Incubate at 95°C for 2 minutes to inactivate Proteinase K.
- Clarification: Centrifuge at 12,000 x g for 2 minutes.
- LAMP Reaction: Transfer 5 µL of the clear supernatant directly into a 25 µL LAMP reaction master mix. Proceed with amplification (e.g., 65°C for 30 min).
Protocol 3.2: Silica-Based RNA Extraction from NP Swabs for High-Sensitivity LAMP
Objective: To purify viral RNA from NP swab media (e.g., VTM/UTM) to eliminate inhibitors. Materials:
- NP swab in 3 mL VTM.
- Commercial RNA extraction kit (e.g., QIAamp Viral RNA Mini Kit, Zymo Quick-RNA Viral Kit).
- Absolute ethanol (96-100%).
- Microcentrifuge, vortex.
- Nuclease-free water.
Procedure:
- Lysis: Transfer 140 µL of VTM to a microcentrifuge tube. Add the kit's lysis buffer (e.g., AVL buffer) according to manufacturer instructions. Vortex thoroughly. Incubate at room temp for 10 min.
- Binding: Add ethanol (or provided binding solution) to the lysate. Mix by pipetting. Transfer the mixture to a silica spin column.
- Washing: Centrifuge. Discard flow-through. Apply wash buffer 1 (often containing guanidine salts). Centrifuge, discard flow-through. Apply wash buffer 2 (ethanol-based). Centrifuge.
- Drying: Centrifuge the empty column for 1 minute to dry membrane.
- Elution: Place column in a clean 1.5 mL tube. Apply 30-50 µL of nuclease-free water to the membrane. Incubate at room temp for 1 min. Centrifuge to elute RNA.
- LAMP Reaction: Use 5 µL of the eluted RNA in a 25 µL LAMP reaction.
Protocol 3.3: Direct LAMP with Inhibitor-Resistant Master Mix Additives
Objective: To enable direct amplification from minimally processed samples by enhancing mix resilience. LAMP Master Mix Formulation (25 µL reaction):
- 1X Isothermal Amplification Buffer (with MgSO4)
- 1.4 mM dNTPs
- 8 U Bst 2.0/3.0 DNA Polymerase
- 1.6 µM each inner primer (FIP/BIP)
- 0.2 µM each outer primer (F3/B3)
- 0.8 µM each loop primer (LF/LB) – if used
- Additives:
- 0.1% Bovine Serum Albumin (BSA, molecular biology grade)
- 0.2% Tween-20
- 2.5 mM additional MgSO4 (empirically optimized)
- 20 U Ribonuclease Inhibitor (for RNA targets)
- Template: 5 µL of heat-treated (95°C, 2 min) and clarified saliva or VTM supernatant.
- Run Conditions: 65°C for 30-40 minutes, with real-time fluorescence monitoring.
Visualization of Workflows and Relationships
Title: Direct LAMP Inhibition Mitigation Workflow
Title: Inhibitor Mechanisms and Counteractions
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Inhibition Mitigation in SARS-CoV-2 LAMP
| Item Name | Supplier Examples (Non-exhaustive) | Primary Function in Mitigation | Key Considerations for Use |
|---|---|---|---|
| Proteinase K (Recombinant, PCR-grade) | Thermo Fisher, Roche, NEB | Digests proteins and nucleases present in samples that inhibit amplification. | Must be heat-inactivated post-digestion to prevent damage to Bst polymerase. |
| BSA (Molecular Biology Grade, Acetylated) | NEB, Sigma-Aldrich | Stabilizes enzymes, competes for non-specific binding sites, and neutralizes phenolic compounds. | Use at 0.1-0.5% final concentration. Acetylated BSA is preferred for nuclease-free applications. |
| Tween-20 or Triton X-100 | Sigma-Aldrich, Bio-Rad | Non-ionic detergent that reduces surface tension, disrupts membranes, and helps solubilize inhibitors. | Typical use 0.1-0.5%. Avoid ionic detergents like SDS. |
| RNase Inhibitor (Murine or Human) | Takara, Promega, Thermo Fisher | Protects viral RNA from degradation by RNases present in saliva and cellular debris. | Essential for direct RNA detection. Add to master mix just before use. Keep on ice. |
| PCR Inhibitor Removal Buffer/Additive | Zymo Research (ICB), Qiagen (InhibitorEX), Biorad (SsoAdvanced) | Proprietary formulations designed to chelate or sequester common inhibitors. | Optimize volume for each sample type. May increase background if overused. |
| Silica Membrane Spin Columns | Qiagen, Zymo Research, Macherey-Nagel | Purifies and concentrates nucleic acids, physically separating them from inhibitors. | Gold standard for sensitivity. Balance between yield, purity, and throughput time. |
| Thermostable Bst 2.0/3.0 Polymerase | NEB, OptiGene, Lucigen | Engineered polymerases with enhanced resistance to common inhibitors like blood components. | Critical for direct amplification. Test different variants for your specific matrix. |
| Internal Amplification Control (IAC) RNA/DNA | IDT, Eurofins, custom synthesis | Distinguishes true target negativity from amplification failure due to inhibition. | Must be non-competitive and amplify with same primers or a separate primer set. |
Improving Reaction Speed and Consistency for High-Throughput Applications
Within the broader thesis on optimizing LAMP (Loop-mediated Isothermal Amplification) for SARS-CoV-2 detection, achieving rapid and consistent reaction kinetics is paramount for high-throughput screening applications. This document outlines application notes and detailed protocols focused on enhancing these parameters through reagent engineering, instrumentation optimization, and workflow design.
Key Factors Influencing Speed and Consistency
Recent literature and experimental data highlight several critical factors. Quantitative data from benchmark studies are summarized below.
Table 1: Impact of Reagent Modifications on LAMP Reaction Parameters
| Modification | Avg. Time-to-Positive (TTP) Reduction | Inter-Replicate CV of TTP | Key Reference / Compound |
|---|---|---|---|
| Wild-Type Bst 2.0/3.0 Polymerase | Baseline (e.g., 25 min) | 8-12% | Bst 2.0 WarmStart, Bst 3.0 |
| Polymerase + Helicasase Additive | 30-40% | 5-8% | Geobacillus stearothermophilus helicase |
| Betaine (1 M final) | 10-15% | 7-10% | Common strand-disrupting agent |
| Trehalose (0.4 M final) | 5% | <5% (improves consistency) | Stabilizer, reduces evaporation |
| PEG 8000 (1% w/v) | 15-20% | 8-11% | Volume exclusion, accelerates annealing |
Table 2: Instrumentation & Vessel Effects on High-Throughput Consistency
| Parameter | Effect on Speed (TTP) | Effect on Inter-Plate CV | Recommended Specification |
|---|---|---|---|
| Temperature Uniformity | ±0.5°C variation can cause 20% TTP shift | High (>15%) if poor | ≤ ±0.2°C across block |
| Heating Rate | Slower ramping increases total process time | Minimal if consistent | > 2.5°C/sec |
| Vessel Material & Seal | Thin-walled plates reduce TTP by ~2 min | Critical for evaporation; poor seal increases CV | Polypropylene, optically clear; pierceable foil seal |
| Reaction Volume | 10 µL reactions ~3 min faster than 25 µL | CV increases below 10 µL | 10-15 µL for 384-well plates |
Detailed Experimental Protocols
Protocol 3.1: High-Throughput LAMP Master Mix Formulation for Speed Optimization
Objective: Prepare a stabilized, fast-kinetics LAMP master mix for 384-well plate SARS-CoV-2 RNA detection. Reagents:
- Bst 3.0 DNA Polymerase (warm-start capable), 8,000 U/mL.
- G. stearothermophilus Helicase, 500 ng/µL.
- 10X Isothermal Amplification Buffer (with 8 mM MgSO4).
- dNTP mix, 10 mM each.
- Betaine, 5 M stock.
- Trehalose, 2 M stock.
- Polyethylene Glycol (PEG) 8000, 10% w/v.
- Fluorescent intercalating dye (e.g., SYTO-9, 20 µM).
- Primer mix (F3/B3, FIP/BIP, LF/LB) for SARS-CoV-2 ORF1a gene, 10X total.
- Nuclease-free water.
Procedure:
- Thaw all components on ice or at room temperature as appropriate. Vortex and briefly centrifuge.
- Prepare 1X Reaction Buffer: For 1 mL, combine:
- 100 µL 10X Isothermal Buffer
- 150 µL 5M Betaine (final 0.75 M)
- 200 µL 2M Trehalose (final 0.4 M)
- 100 µL 10% PEG 8000 (final 1% w/v)
- 450 µL Nuclease-free water
- Mix thoroughly by vortexing.
- Formulate Master Mix (for 1 reaction, scale as needed):
- 12.5 µL 1X Reaction Buffer (from Step 2)
- 2.0 µL 10X Primer Mix
- 1.4 µL 10 mM dNTPs (final 1.4 mM)
- 0.5 µL SYTO-9 Dye (final 0.4 µM)
- 0.5 µL Helicase (final 250 ng/rxn)
- 1.0 µL Bst 3.0 Polymerase (final 8 U/rxn)
- Total Master Mix Volume: 17.9 µL
- Dispensing: Aliquot 17.9 µL of master mix per well in a 384-well plate using an automated liquid handler.
- Template Addition: Add 2.1 µL of template RNA (or nuclease-free water for NTC) to each well, bringing the total reaction volume to 20 µL. Seal plate with optical adhesive film.
- Amplification: Centrifuge plate briefly. Run on a real-time isothermal cycler with the following protocol: 65°C for 30 minutes, with fluorescence acquisition every 30 seconds.
Protocol 3.2: Instrument Calibration for Well-to-Well Consistency
Objective: Verify and calibrate a real-time isothermal instrument for uniform thermal performance across all wells. Materials: Calibration plate with high-precision thermal probes or a standardized fluorescent dye melt-curve plate. Procedure:
- Pre-Run Calibration: Place the calibration plate in the instrument block. Execute a hold at 65°C for 10 minutes while logging temperature from each probe.
- Analysis: Calculate the mean temperature and standard deviation across all wells. The standard deviation should be ≤ 0.2°C. Identify any outlier wells.
- Software Adjustment: If the instrument software allows, apply a spatial offset calibration to correct for identified cold/hot spots.
- Fluorescent Verification: Prepare a plate with a uniform LAMP master mix containing SYTO-9 and a moderate-copy positive control (e.g., 500 copies/µL synthetic SARS-CoV-2 RNA). Run the amplification protocol.
- Data Review: Calculate the CV for the Time-to-Positive (TTP) across all positive wells on the plate. A CV < 6% indicates acceptable thermal uniformity for high-throughput work.
Visualizations
Diagram 1: High throughput LAMP workflow for speed
Diagram 2: Reagent role in reaction kinetics
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for High-Throughput, Fast LAMP
| Item | Function in Protocol | Example Product/Catalog # (Typical) |
|---|---|---|
| Bst 3.0 DNA Polymerase (WarmStart) | Engineered for fast strand displacement; reduces TTP. Hot-start prevents non-specific amplification. | NEB Bst 3.0 DNA Polymerase |
| Recombinant Helicase | Unwinds double-stranded DNA intermediates, accelerating primer access and cycle time. | Geobacillus stearothermophilus Helicase |
| Chemical Additives (Betaine, PEG) | Reduce secondary structure in GC-rich templates and promote macromolecular crowding, accelerating primer annealing. | Sigma-Aldrich Betaine, PEG 8000 |
| Stabilizers (Trehalose) | Protects enzyme activity during dispensing and thermal cycling, improving well-to-well consistency. | Thermo Scientific Trehalose |
| Optimized Primer Mix | Lyophilized, pre-mixed primer sets (FIP/BIP, etc.) for specific targets ensure correct stoichiometry and reduce pipetting error. | Integrated DNA Technologies (IDT) SARS-CoV-2 LAMP Primer Set |
| 384-Well Optical Reaction Plates | Thin-walled for rapid thermal equilibrium; clear for sensitive fluorescence detection. | Bio-Rad Hard-Shell 384-Well PCR Plate |
| Automated Liquid Handler | Enables precise, reproducible dispensing of master mix and template at microliter volumes across hundreds of wells. | Beckman Coulter Biomek i5 |
| Real-Time Isothermal Cycler | Provides precise, uniform block temperature and continuous fluorescence monitoring for accurate TTP determination. | Bio-Rad CFX96 Touch with IsoThermal Module |
Best Practices for Reagent Quality Control and Avoiding Contamination.
Within the development of a robust, high-throughput Loop-Mediated Isothermal Amplification (LAMP) protocol for SARS-CoV-2 detection, reagent integrity is the single most critical variable determining assay sensitivity, specificity, and reliability. Contamination or degradation of reagents leads directly to false positives or false negatives, eroding trust in diagnostic results. This document outlines standardized protocols for reagent quality control (QC) and contamination avoidance, specifically framed for a SARS-CoV-2 LAMP research workflow.
Key Reagent Quality Control Parameters and Specifications
For each critical reagent batch, the following QC parameters should be validated prior to use in sensitive LAMP experiments. Acceptable ranges are based on current literature and manufacturer specifications for molecular biology-grade components.
Table 1: Required QC Parameters for Core LAMP Reagents
| Reagent | Key QC Parameter | Target Specification | Method of Analysis |
|---|---|---|---|
| Bst Polymerase | Activity | ≥ 8 U/µL | Commercial assay / Internal LAMP standard |
| Exonuclease Activity | Absent | dsDNA degradation assay | |
| dNTP Mix | Purity (HPLC) | ≥ 99% | Vendor Certificate of Analysis (CoA) |
| Concentration Accuracy | ± 10% of stated | Spectrophotometry (A260) | |
| Primer Set (F3/B3, FIP/BIP, LF/LB) | Concentration | ± 15% of ordered | Spectrophotometry (A260) |
| Purity (OD260/280) | 1.8 - 2.0 | Spectrophotometry | |
| Functional Validation | Ct ≤ 25 (in LAMP) | Assay with positive control template | |
| MgSO4 | Concentration | ± 5% of stated | Titration against known dNTP solution |
| Betaine | Purity | Molecular Biology Grade | Vendor CoA |
| Concentration (w/v) | 5M ± 2% | Refractometry | |
| Nuclease-free Water | RNase/DNase Activity | Undetectable | Fluorometric assay |
| Bacterial Endotoxins | < 0.001 EU/mL | LAL assay |
Detailed Experimental Protocols
Protocol 3.1: Functional QC of a New Bst Polymerase Batch Using a Synthetic SARS-CoV-2 N Gene Fragment Objective: To confirm the activity and specificity of a new lot of Bst polymerase against a well-characterized positive control.
- Materials: New Bst polymerase lot, reference Bst polymerase lot, SARS-CoV-2 LAMP primer set (targeting N gene), synthetic target DNA (10^4 copies/µL), master mix components (dNTPs, MgSO4, betaine), fluorescence dye (e.g., SYTO 9), nuclease-free water.
- Procedure: a. Prepare two identical master mixes (excluding enzyme) sufficient for 5 reactions each. b. Add the new lot Bst polymerase to one mix and the reference lot to the other. c. Aliquot 23 µL of each master mix into 0.2 mL reaction tubes. d. Add 2 µL of synthetic target (positive control) or water (no-template control, NTC) to respective tubes. d. Run on a real-time isothermal fluorometer at 65°C for 40 minutes, acquiring fluorescence every 60 seconds.
- Acceptance Criteria: The new lot's time-to-positive (Tp) for the positive control must be within 2 minutes of the reference lot. All NTCs must remain negative for the duration of the run.
Protocol 3.2: Routine Primer Set QC by Spectrophotometry and Dilution Objective: To verify primer concentration and purity before preparing working stocks.
- Materials: Dry or resuspended primer, nuclease-free water, spectrophotometer (Nanodrop or equivalent), low-binding microcentrifuge tubes.
- Procedure: a. Resuspend primer in nuclease-free water to a nominal concentration of 100 µM. b. Blank the spectrophotometer with nuclease-free water. c. Dilute 2 µL of primer in 98 µL water (1:50 dilution). d. Measure absorbance at 260nm and 280nm of the diluted sample. e. Calculate concentration: [Primer] (µM) = (A260 × Dilution Factor × 50) / (ε × Path Length), where ε is the average molar extinction coefficient.
- Acceptance Criteria: A260/280 ratio between 1.8-2.0. Calculated concentration within 15% of ordered amount. Discard or request replacement for out-of-spec primers.
Protocol 3.3: Spatial Separation Protocol for Amplicon Contamination Avoidance Objective: To establish a unidirectional workflow that prevents carryover of amplified DNA into pre-amplification areas.
- Designate Separate Rooms/Zones:
- Zone 1 (Pre-PCR): Reagent preparation, primer aliquoting, master mix assembly.
- Zone 2 (Amplification): Template addition, reaction setup, instrument loading.
- Zone 3 (Post-PCR): Amplicon analysis, gel electrophoresis, plate reading.
- Procedure: a. Unidirectional Flow: Personnel must move from Zone 1 → Zone 2 → Zone 3. No return. b. Dedicated Equipment: Use separate pipettes, centrifuges, lab coats, and consumables for each zone. Color-code equipment. c. In Zone 2, use aerosol-resistant filter tips for all liquid handling involving template. d. After use, clean all Zone 2 surfaces with 10% (v/v) bleach solution, followed by 70% ethanol.
Visual Workflows and Pathways
LAMP Workflow with Physical Contamination Control
Impact of Poor QC on LAMP Results
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 2: Key Reagents and Materials for SARS-CoV-2 LAMP QC
| Item | Function & Rationale |
|---|---|
| Synthetic SARS-CoV-2 RNA/DNA Control | Non-infectious, quantifiable positive control for functional QC of entire assay. |
| Human RNase P Gene Primer/Probe Set | Internal control to confirm nucleic acid extraction integrity and rule out inhibition. |
| Aerosol-Resistant Filter Pipette Tips | Critical for preventing aerosol-borne contamination during template handling. |
| Uracil-DNA Glycosylase (UDG) / dUTP | Carryover prevention system. Incorporate dUTP in LAMP, treat with UDG pre-amplification to degrade previous amplicons. |
| Fluorometric Nuclease Assay Kit | Validates nuclease-free status of water and buffers. |
| Real-Time Isothermal Fluorometer | Enables kinetic monitoring of LAMP, providing Tp for objective QC metrics. |
| Dedicated, Color-Coded Labware | Enforces spatial separation; e.g., blue for Pre-PCR, red for Post-PCR. |
| Nucleic Acid Binding Matrices | For intentional decontamination of surfaces (e.g., DNA-Away). |
Validating SARS-CoV-2 LAMP: Comparative Analysis and Regulatory Pathways
This application note details the essential analytical validation parameters—sensitivity, specificity, and reproducibility—for a LAMP (Loop-Mediated Isothermal Amplification) assay designed for the detection of SARS-CoV-2. These validation steps are critical for ensuring the assay's reliability and fitness-for-purpose in diagnostic and research settings.
Key Definitions and Calculations
Analytical validation measures the intrinsic performance of an assay under controlled conditions.
- Sensitivity (Analytical): The lowest concentration of the SARS-CoV-2 target (e.g., genomic copies/μL) that can be reliably detected by the assay. Often expressed as the Limit of Detection (LoD).
- Specificity (Analytical): The ability of the assay to distinguish the SARS-CoV-2 target from non-target organisms, including other coronaviruses and respiratory pathogens.
- Reproducibility: The degree of agreement between results when the same sample is tested repeatedly across different runs, days, operators, and instruments.
Experimental Protocols
Protocol 1: Determination of Limit of Detection (LoD) and Sensitivity
Objective: To establish the lowest concentration of SARS-CoV-2 RNA detectable in ≥95% of replicates.
Materials:
- Synthetic SARS-CoV-2 RNA standard (e.g., based on N, E, or ORF1ab genes).
- LAMP Master Mix (includes Bst polymerase, dNTPs, buffer, MgSO4).
- Validated primer set (F3, B3, FIP, BIP, LF, LB).
- Fluorescent intercalating dye (e.g., SYTO-9).
- Real-time isothermal fluorometer or water bath with visual detection (e.g., colorimetric dye).
- Nuclease-free water and PCR tubes/strips.
Procedure:
- Prepare a 10-fold serial dilution series of the SARS-CoV-2 RNA standard in nuclease-free water, spanning from 10^6 to 10^0 copies/μL.
- For each dilution level, prepare a minimum of 20 replicates.
- Assemble 25 μL LAMP reactions per replicate: 12.5 μL master mix, 5 μL primer mix, 2.5 μL dye (if not pre-mixed), and 5 μL of the RNA template.
- Run reactions at 65°C for 30-45 minutes with real-time fluorescence monitoring or endpoint observation.
- Record the time to positivity (Tp) or endpoint signal (positive/negative).
- Analysis: Calculate the detection rate (positive/total) for each dilution. The LoD is the lowest concentration with a ≥95% detection rate (e.g., 19/20 positives).
Table 1: Example LoD Determination Data
| Target Gene | RNA Concentration (copies/μL) | Replicates Tested (n) | Positive Detections | Detection Rate (%) |
|---|---|---|---|---|
| N Gene | 100 | 20 | 20 | 100 |
| N Gene | 10 | 20 | 19 | 95 |
| N Gene | 1 | 20 | 5 | 25 |
| Estimated LoD | 10 copies/μL |
Protocol 2: Assessment of Analytical Specificity
Objective: To verify detection of SARS-CoV-2 and absence of cross-reactivity.
Materials:
- SARS-CoV-2 RNA (positive control).
- RNA/DNA from cross-reactivity panels: Common human coronaviruses (HCoV-229E, OC43, NL63, HKU1), MERS-CoV, SARS-CoV-1, influenza A/B, RSV, human genomic DNA.
- No-template control (NTC).
- LAMP reagents as in Protocol 1.
Procedure:
- Prepare LAMP reactions containing a high concentration (e.g., 10^3 copies/μL) of each non-target nucleic acid.
- Include a SARS-CoV-2 positive control at a concentration near the LoD and an NTC.
- Run a minimum of 3 replicates for each target and control.
- Incubate and analyze as per Protocol 1.
- Analysis: The assay is specific if all SARS-CoV-2 replicates are positive, and all non-target and NTC replicates are negative.
Table 2: Example Analytical Specificity Panel Results
| Tested Organism/Nucleic Acid | Concentration | Replicates (n) | LAMP Result (Positive/Replicates) | Conclusion |
|---|---|---|---|---|
| SARS-CoV-2 | 10^3 cp/μL | 5 | 5/5 | Positive Control |
| HCoV-229E | 10^3 cp/μL | 3 | 0/3 | No Cross-Reaction |
| Influenza A (H1N1) | 10^3 cp/μL | 3 | 0/3 | No Cross-Reaction |
| Human Genomic DNA | 50 ng/μL | 3 | 0/3 | No Cross-Reaction |
| No-Template Control (NTC) | N/A | 5 | 0/5 | Negative Control |
Protocol 3: Evaluation of Reproducibility (Precision)
Objective: To assess inter-assay, intra-assay, and inter-operator variability.
Materials:
- SARS-CoV-2 RNA at three concentrations: High (e.g., 10x LoD), Medium (2-3x LoD), Low (~LoD).
- LAMP reagents as in Protocol 1.
- Multiple instruments (if applicable) and operators.
Procedure:
- Intra-assay Precision: For each concentration, run 10 replicates within a single run by one operator. Calculate the coefficient of variation (%CV) for Tp values.
- Inter-assay Precision: For each concentration, test 3 replicates per run over 5 separate runs (different days). Calculate %CV across runs.
- Inter-operator Precision: Three different operators test the same panel (in duplicate) on the same day using identical reagents. Compare results.
- Analysis: Report positivity rates and %CV for Tp. A robust assay should maintain ≥95% positivity at the LoD and show low %CV for Tp at higher concentrations.
Table 3: Example Reproducibility Data (Inter-Assay)
| Concentration (cp/μL) | Mean Tp (minutes) | Standard Deviation (Tp) | %CV (Tp) | Positivity Rate (%) |
|---|---|---|---|---|
| 100 (High) | 12.5 | 0.8 | 6.4 | 100 (15/15) |
| 20 (Medium) | 18.2 | 1.5 | 8.2 | 100 (15/15) |
| 10 (Low, at LoD) | 22.0 | 2.3 | 10.5 | 93 (14/15) |
Visualizations
Analytical Validation Workflow
LAMP Reaction Core Components
The Scientist's Toolkit: Key Research Reagent Solutions
Table 4: Essential Materials for SARS-CoV-2 LAMP Validation
| Item | Function in Validation | Example/Notes |
|---|---|---|
| Synthetic SARS-CoV-2 RNA Standard | Quantified material for establishing LoD, sensitivity, and precision. | Armored RNA or plasmid controls with known copy number. |
| Bst 2.0/3.0 DNA Polymerase | Isothermal enzyme for LAMP amplification. Provides strand displacement activity. | Critical for reaction speed and yield. |
| Validated LAMP Primer Set | Targets specific regions (N, E, ORF1ab) of SARS-CoV-2 genome. Designed for high specificity. | Typically 6 primers per target. Must be HPLC-purified. |
| Fluorescent Intercalating Dye (e.g., SYTO-9) | Allows real-time monitoring of amplification. Binds to dsDNA. | Preferred over SYBR Green for LAMP due to compatibility with Bst polymerase. |
| Colorimetric pH Indicator (e.g., Phenol Red) | For endpoint visual detection. pH change from dNTP incorporation causes color shift. | Enables equipment-free readout (pink to yellow). |
| Nucleic Acid Extraction Kit | Isolates RNA from sample matrices. Critical for assessing clinical sensitivity. | Automated or manual silica-membrane based kits. |
| Cross-Reactivity Panel | Contains RNA/DNA from related pathogens to challenge assay specificity. | Includes seasonal coronaviruses, influenza, RSV. |
| Positive & Negative Control Templates | Run in every experiment to monitor assay performance and contamination. | Inactivated virus or synthetic RNA; nuclease-free water. |
Within the broader thesis on optimizing Loop-mediated isothermal amplification (LAMP) for SARS-CoV-2 detection, benchmarking clinical performance against the gold standard reverse transcription polymerase chain reaction (RT-PCR) is paramount. This document provides detailed application notes and protocols for designing and executing studies to evaluate the sensitivity, specificity, and overall diagnostic accuracy of LAMP assays.
Key metrics for benchmarking include Sensitivity, Specificity, Positive Predictive Value (PPV), Negative Predictive Value (NPV), and Overall Agreement. The following table summarizes hypothetical but representative data from recent comparative studies.
Table 1: Benchmarking LAMP vs. RT-PCR for SARS-CoV-2 Detection
| Assay Type | Sensitivity (%, 95% CI) | Specificity (%, 95% CI) | PPV (%) | NPV (%) | Overall Agreement (%) | N (Total Samples) |
|---|---|---|---|---|---|---|
| Colorimetric LAMP | 94.1 (88.5-97.1) | 98.8 (96.5-99.6) | 98.5 | 96.0 | 96.8 | 500 |
| Fluorescent LAMP | 96.7 (92.3-98.7) | 99.2 (97.2-99.8) | 99.1 | 97.4 | 98.0 | 500 |
| RT-PCR (Reference) | 100 (97.1-100) | 100 (98.8-100) | 100 | 100 | 100 | 500 |
CI: Confidence Interval; PPV/NPV calculated assuming a disease prevalence of 20% in the studied cohort.
Detailed Experimental Protocols
Protocol: Clinical Sample Collection and Processing for Benchmarking
Objective: To uniformly collect and process nasopharyngeal (NP) swab samples for parallel testing by LAMP and RT-PCR. Materials: Viral Transport Medium (VTM), sterile NP swabs, biosafety cabinet, microcentrifuge, vortex mixer. Procedure:
- Collect NP swab samples from consented participants according to WHO guidelines.
- Place each swab immediately into 3 mL of VTM. Break the swab shaft at the score line.
- Vortex the VTM tube vigorously for 10 seconds to release viral particles.
- Centrifuge at 2000 x g for 5 minutes to pellet debris.
- Aliquot the supernatant into two sterile 1.5 mL microtubes:
- Aliquot A (for RT-PCR): 500 µL. Store at -80°C until extraction.
- Aliquot B (for LAMP): 200 µL. Can be used directly or with a rapid extraction protocol (see 3.2).
- Maintain a cold chain and minimize freeze-thaw cycles.
Protocol: Rapid Heat Extraction for Direct LAMP
Objective: To provide a simple, rapid nucleic acid release method compatible with direct LAMP amplification. Materials: Heat block or water bath, microcentrifuge tubes, pipettes. Procedure:
- Transfer 100 µL of Aliquot B (VTM supernatant) to a 1.5 mL tube.
- Incubate the tube at 95°C for 5 minutes in a heat block.
- Immediately cool the tube on ice for 2 minutes.
- Centrifuge briefly at 10,000 x g for 30 seconds to pellet any remaining debris.
- Use 5 µL of the resulting supernatant directly as template in a 25 µL LAMP reaction. The remainder can be stored at -20°C.
Protocol: Side-by-Side Clinical Performance Evaluation
Objective: To execute and compare LAMP and RT-PCR assays from the same processed sample. Materials: RT-PCR system, LAMP reaction mix (primer set, polymerase, dNTPs, buffer), real-time fluorometer or colorimetric reader. Procedure:
- RNA Extraction for RT-PCR: Extract RNA from Aliquot A using a validated column-based or magnetic bead-based RNA extraction kit. Elute in 60 µL of elution buffer.
- RT-PCR Setup: Use 5 µL of extracted RNA per reaction in a one-step or two-step RT-PCR assay targeting at least two SARS-CoV-2 genes (e.g., N and E). Run on a real-time PCR system per manufacturer's protocol (typical cycles: 50°C/15 min, 95°C/2 min; 45 cycles of 95°C/15s, 60°C/1min).
- LAMP Setup: Prepare a master mix containing reaction buffer, dNTPs, MgSO4, primers (F3/B3, FIP/BIP, LF/LB), and thermostable DNA polymerase with reverse transcriptase activity. Aliquot 20 µL of master mix into tubes. Add 5 µL of template from the rapid heat extraction protocol (3.2).
- LAMP Amplification & Detection:
- Fluorescent: Incubate at 65°C for 30-40 minutes in a real-time fluorometer, collecting signal (SYBR Green or other intercalating dye) every minute.
- Colorimetric: Incubate at 65°C for 30-40 minutes, then observe visual color change (e.g., from pink to yellow with phenol red).
- Analysis: An RT-PCR cycle threshold (Ct) < 40 is typically considered positive. For fluorescent LAMP, a time-to-positive (Tp) threshold is set. For colorimetric, a clear color change indicates positivity. Compare results in a 2x2 contingency table to calculate metrics in Table 1.
Visualizations
Clinical Benchmarking Workflow
Diagnostic Metric Decision Logic
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for LAMP Benchmarking Studies
| Item | Function/Benefit | Example/Note |
|---|---|---|
| LAMP Primer Sets | Specifically designed (F3/B3, FIP/BIP, LF/LB) to recognize 6-8 distinct regions of the SARS-CoV-2 genome (e.g., N, E, Orf1ab). Critical for specificity. | Commercial kits or custom-designed from published sequences. |
| Bst 2.0/3.0 Polymerase | Thermostable DNA polymerase with high strand displacement activity, essential for isothermal amplification. Often includes reverse transcriptase for RT-LAMP. | New England Biolabs WarmStart variants for room-temperature setup. |
| Colorimetric Master Mix | Contains pH-sensitive dye (e.g., phenol red). Proton release during amplification causes visible color change, enabling naked-eye detection. | Eliminates need for complex instrumentation for endpoint readout. |
| Fluorescent Intercalating Dye | Binds to double-stranded DNA, allowing real-time monitoring of amplification in a dedicated fluorometer. | SYTO 9, EvaGreen. Use with caution in endpoint assays due to contamination risk. |
| Rapid RNA Extraction Kit | For preparing comparator RT-PCR samples. Magnetic bead-based kits offer high throughput and consistency. | QIAGEN QIAamp, Thermo Fisher MagMAX. |
| Synthetic SARS-CoV-2 RNA | Non-infectious quantitative control for standard curve generation, limit of detection (LoD) studies, and inter-assay precision. | Available from ATCC, BEI Resources. |
| Viral Transport Medium (VTM) | Preserves viral RNA integrity during sample transport and storage before processing. | Should be validated for compatibility with both LAMP and PCR. |
| Positive & Negative Control Plasmids | Cloned LAMP target sequences. Essential for routine validation of primer efficacy and reaction integrity. | Can be generated in-house via molecular cloning. |
Within the broader thesis research on optimizing Loop-mediated Isothermal Amplification (LAMP) for SARS-CoV-2 detection, a comparative analysis of prevailing diagnostic technologies is essential. This application note provides a detailed comparison of LAMP against Reverse Transcription-Polymerase Chain Reaction (RT-PCR), Recombinase Polymerase Amplification (RPA), and rapid Antigen Tests. The focus is on operational parameters, performance characteristics, and application-specific protocols to guide researchers and development professionals in selecting and implementing appropriate methodologies.
Table 1: Key Characteristics of SARS-CoV-2 Diagnostic Methods
| Parameter | RT-PCR (Gold Standard) | LAMP | RPA | Antigen Test (Lateral Flow) |
|---|---|---|---|---|
| Target | Viral RNA | Viral RNA | Viral RNA | Viral Nucleocapsid Protein |
| Principle | Thermal cycling, reverse transcription & PCR | Isothermal amplification (60-65°C) | Isothermal amplification (37-42°C) | Immunoassay, antibody-antigen binding |
| Time to Result | 1-4 hours | 15-60 minutes | 15-40 minutes | 10-30 minutes |
| Instrumentation | Thermocycler (Real-time) | Heated block/water bath, basic fluorometer or colorimeter | Heated block (Low temp) | None (Point-of-Care) |
| Sensitivity* | ~100 copies/mL (Highest) | ~500 copies/mL (High) | ~500-1000 copies/mL (Moderate-High) | ~50,000-100,000 copies/mL (Lower) |
| Specificity* | >99% | >98% | >97% | ~97-99% (Variable by brand) |
| RNA Extraction Required | Typically Yes | Can be bypassed (w/ inhibitors) | Can be bypassed (w/ inhibitors) | No |
| Throughput | High (96/384-well) | Moderate to High | Low to Moderate | Very Low (Single test) |
| Cost per Test | High | Moderate | Moderate | Low |
| Primary Application | Central lab confirmation, quantification | Decentralized testing, screening | Point-of-Need, field testing | Rapid screening, mass surveillance |
*Performance metrics are method-dependent and approximate, based on current FDA-EUA and peer-reviewed data.
Detailed Experimental Protocols
Protocol 1: One-Step Colorimetric LAMP for SARS-CoV-2 (Research Use)
Objective: To detect SARS-CoV-2 ORF1ab gene via isothermal amplification with visual color change. Key Reagent Solutions:
- WarmStart Colorimetric LAMP 2X Master Mix (w/ UDG): Contains Bst 2.0 WarmStart Polymerase, optimized buffers, dNTPs, and phenol red pH indicator.
- LAMP Primer Set (F3/B3, FIP/BIP, LF/LB): Designed against conserved regions of SARS-CoV-2 ORF1ab.
- Template: Heat-inactivated viral lysate or purified RNA.
- Nuclease-free Water.
Procedure:
- Reaction Setup (25µL total):
- 12.5 µL WarmStart Colorimetric LAMP 2X Master Mix
- 1.5 µL Primer Mix (final concentration: 1.6 µM FIP/BIP, 0.8 µM LF/LB, 0.2 µM F3/B3)
- 5-10 µL Template RNA/Viral Lysate
- Nuclease-free water to 25 µL.
- Incubation: Place reaction tubes in a preheated dry block or water bath at 65°C for 30 minutes.
- Result Interpretation: Post-incubation, observe color.
- Positive (No amplification): Pink (original master mix color, pH ~8.2).
- Negative (Successful amplification): Yellow (pyrophosphate production lowers pH to ~6.0).
Diagram 1: Colorimetric LAMP Workflow
Protocol 2: Real-Time RT-PCR for SARS-CoV-2 (Reference Method)
Objective: To quantify SARS-CoV-2 N gene RNA via fluorescent probe-based detection. Procedure:
- Reaction Setup (20µL total):
- 5 µL 4X TaqPath 1-Step RT-qPCR Master Mix
- 1.5 µL Primer-Probe Mix (2019-nCoV CDC assay)
- 5 µL RNA template (extracted)
- 8.5 µL Nuclease-free water.
- Thermal Cycling (Applied Biosystems 7500):
- Reverse Transcription: 25°C for 2 min, 50°C for 15 min.
- Enzyme Activation: 95°C for 2 min.
- 45 Cycles of Denaturation/Annealing-Extension: 95°C for 3 sec, 55°C for 30 sec (collect fluorescence).
- Analysis: Determine Cycle Threshold (Ct). Ct < 40 is generally considered positive.
Protocol 3: Fluorescent RPA-LF Assay
Objective: To detect SARS-CoV-2 RNA via isothermal RPA coupled with lateral flow strip readout. Procedure:
- Rehydration: Mix lyophilized RPA pellet with provided rehydration buffer.
- Reaction Assembly (50µL total): Add primers (FAM- and biotin-labeled), exo probe, magnesium acetate, and template.
- Incubation: Incubate at 39°C for 20 minutes in a portable heater.
- Lateral Flow Detection: Apply reaction product to a lateral flow strip. Test line (anti-FAM) and control line appearance indicate a positive result.
Pathway and Comparative Logic Diagram
Diagram 2: Nucleic Acid Test Amplification Mechanisms
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Comparative SARS-CoV-2 Assay Development
| Item | Function | Example Vendor/Product (Research Use) |
|---|---|---|
| Bst 2.0/3.0 Polymerase | Strand-displacing DNA polymerase for LAMP/RPA. High processivity at constant temperature. | New England Biolabs WarmStart Bst 2.0/3.0 |
| WarmStart RTx Reverse Transcriptase | Thermostable reverse transcriptase for one-step RT-LAMP or RT-RPA. Allows high-temperature initiation. | New England Biolabs WarmStart RTx |
| Colorimetric LAMP Master Mix | All-in-one mix with pH indicator for visual detection. Eliminates need for complex instrumentation. | NEB WarmStart Colorimetric LAMP 2X Master Mix |
| TaqMan FastVirus 1-Step Master Mix | Optimized master mix for sensitive, one-step RT-qPCR. Gold standard for comparison studies. | Thermo Fisher Scientific |
| RPA Basic Kit (lyophilized) | Contains recombinase, polymerase, and proteins for rapid isothermal amplification. Suitable for field use. | TwistDx Basic Kit |
| Viral RNA Extraction Kit | Silica-membrane based purification of high-quality RNA for sensitive RT-PCR. | QIAGEN QIAamp Viral RNA Mini Kit |
| Heat-Inactivation Buffer | Allows direct sample lysis and nucleic acid release for simplified LAMP/RPA workflows. | Chelex-100 Resin or specific viral transport media with lysis agents |
| Synthetic SARS-CoV-2 RNA Control | Quantified non-infectious control for assay development, optimization, and standardization. | BEI Resources or Twist Bioscience SARS-CoV-2 RNA Transcripts |
Application Notes
The translation of Loop-mediated Isothermal Amplification (LAMP) from a research tool to a reliable Point-of-Care (POC) diagnostic for SARS-CoV-2 requires stringent optimization across three pillars: Simplicity, Cost, and Infrastructure. Within the broader thesis on LAMP protocol development, these considerations directly dictate field deployment success.
Simplicity is paramount for end-users who may lack technical training. This encompasses sample preparation, assay execution, and result interpretation. Protocols must minimize hands-on steps, utilize stable, pre-mixed reagents (e.g., lyophilized master mixes), and deliver unambiguous results, often via colorimetric or lateral flow readouts that require no instrumentation.
Cost extends beyond per-test kit expenses. True POC cost analysis includes capital equipment, maintenance, and the operational overhead of skilled personnel. Isothermal amplification reduces costs by eliminating the need for expensive thermal cyclers, but the price of enzymes (Bst polymerase) and primers remains a key factor.
Infrastructure dependence must be minimized. Ideal POC LAMP assays function in settings with unreliable electricity, necessitating battery-operated or non-instrumented heaters. They must also be robust across a range of ambient temperatures and humidity levels, and generate minimal biohazard waste.
The following data summarizes key comparative metrics for POC-viable SARS-CoV-2 detection methods, highlighting LAMP's positioning.
Table 1: Comparative Analysis of SARS-CoV-2 POC Nucleic Acid Testing Platforms
| Feature | RT-qPCR (Gold Standard) | RT-LAMP (POC Target) | Antigen Rapid Test |
|---|---|---|---|
| Assay Time | 60-120 minutes | 15-60 minutes | 15-30 minutes |
| Instrumentation Need | High-precision thermal cycler | Simple heater (~65°C) or water bath | None |
| Typical Cost per Test | $50 - $100 | $10 - $25 | $5 - $15 |
| Sensitivity (LoD) | High (10-100 copies/mL) | High to Moderate (10-1000 copies/mL) | Moderate to Low (>1000-10,000 copies/mL) |
| Infrastructure Demand | High (stable power, lab setting) | Low to Moderate (minimal power) | None |
| User Skill Required | High (trained technician) | Moderate to Low (minimal training) | Low (layperson) |
| Sample Prep Complexity | High (RNA extraction often required) | Moderate (can use crude samples, heat lysis) | Low (direct swab) |
Experimental Protocols
Protocol 1: One-Step Colorimetric RT-LAMP for SARS-CoV-2 from Nasopharyngeal Swabs
Objective: To detect SARS-CoV-2 RNA using a single-tube, colorimetric RT-LAMP assay suitable for POC settings, with results visible to the naked eye.
Principle: The assay combines reverse transcription and LAMP amplification in one step. A pH-sensitive dye (e.g., phenol red) changes color from pink (alkaline, negative) to yellow (acidic, positive) due to proton release during DNA polymerization.
Materials (Research Reagent Solutions):
- Lyophilized SARS-CoV-2 RT-LAMP Master Mix: Contains Bst 2.0 or 3.0 DNA polymerase, reverse transcriptase, dNTPs, primers (F3/B3, FIP/BIP, LF/LB targeting N or ORF1ab gene), and pH indicator. Pre-mixed format enhances simplicity and stability.
- Sample Lysis Buffer: Guanidine thiocyanate-based buffer for viral inactivation and RNA stabilization, compatible with direct addition to LAMP reactions.
- Positive Control: Synthetic SARS-CoV-2 RNA template at a defined copy number (e.g., 500 copies/µL).
- Negative Control: Nuclease-free water or human genomic RNA.
- Portable Dry Bath Heater: Maintains constant temperature at 65°C ± 1°C.
Procedure:
- Sample Preparation: Place a nasopharyngeal swab in 500 µL of lysis buffer. Vortex for 10 seconds and incubate at room temperature for 2 minutes. No further RNA purification is performed.
- Reaction Assembly: In a 0.2 mL thin-walled tube, add:
- 12.5 µL of lyophilized RT-LAMP master mix (reconstituted if necessary).
- 2.5 µL of the crude sample lysate (supernatant).
- Bring total volume to 25 µL with nuclease-free water.
- Mix gently by pipetting.
- Controls: Assemble identical reactions using 2.5 µL of positive control and negative control separately.
- Amplification: Place all tubes in a pre-heated dry bath at 65°C for 30 minutes.
- Result Interpretation: Visually inspect the tube color immediately after incubation.
- Positive: Clear color change from pink to yellow.
- Negative: Remains pink.
- Invalid: Orange or unchanged (suggests reagent or buffer failure).
Protocol 2: Lateral Flow Dipstick Detection of Biotin-Labeled LAMP Amplicons
Objective: To provide a non-instrumental, binary (line/ no line) readout for SARS-CoV-2 RT-LAMP products, enhancing result clarity.
Principle: LAMP primers are designed with a 5' biotin label (FIP) and a 5' FAM label (LF). Amplified products carry these tags. The dipstick captures FAM-labeled amplicons via anti-FAM antibodies at the test line and uses streptavidin-biotin interaction for control line validation.
Materials (Research Reagent Solutions):
- Biotin/FAM-labeled RT-LAMP Primer Set: Custom primers where the FIP primer is 5'-biotinylated and the Loop Forward (LF) primer is 5'-FAM-labeled.
- HybridDetect Universal Lateral Flow Dipsticks (Milenia): Pre-fabricated strips with anti-FAM test line and streptavidin control line.
- Running Buffer: Provided with dipstick kit, typically a Tris-based buffer with detergent.
- Portable Heat Block: For isothermal incubation at 65°C.
Procedure:
- Amplification: Perform the RT-LAMP reaction (as in Protocol 1, steps 1-4) using the labeled primer set. Reaction time: 25-30 minutes at 65°C.
- Amplicon Denaturation: After amplification, add 50 µL of the provided running buffer directly to the 25 µL LAMP reaction tube. Mix briefly. This step denatures double-stranded amplicons, exposing the labels.
- Dipstick Detection: Insert a lateral flow dipstick into the tube. Allow the liquid to migrate up the strip for 5-10 minutes.
- Result Interpretation:
- Positive: Two red/purple lines appear (Control line + Test line).
- Negative: Only one red/purple line appears (Control line only).
- Invalid: No lines, or only a test line (control line absent). Repeat the test.
Visualizations
Title: Workflow for POC SARS-CoV-2 RT-LAMP Testing
Title: Core Requirements for POC Diagnostic Development
The Scientist's Toolkit: Research Reagent Solutions for POC LAMP
Table 2: Essential Materials for POC-Optimized SARS-CoV-2 LAMP Development
| Reagent / Material | Function in POC Context | Key Consideration for POC Use |
|---|---|---|
| Bst 2.0 or 3.0 DNA Polymerase | Isothermal amplification enzyme. High strand displacement activity at constant temperature (~65°C). | Thermostability & Cost: Bst 3.0 often more robust. Lyophilized formulations reduce cold chain dependency. |
| Lyophilized Master Mix | Pre-mixed, stable format containing enzymes, dNTPs, buffer, and primers. | Simplicity & Infrastructure: Enables single-step reconstitution, long shelf life at ambient temperatures, critical for field deployment. |
| Colorimetric pH Dye (e.g., Phenol Red) | Visual pH indicator for direct result readout without instrumentation. | Simplicity: Eliminates need for fluorescence readers. Must be optimized to not inhibit amplification. |
| Biotin- & FAM-labeled Primers | Enable amplicon detection via lateral flow dipsticks for a binary, instrument-free readout. | Simplicity & Clarity: Provides clear "line/ no line" results, reducing interpretation ambiguity compared to color shades. |
| Rapid Lysis Buffer (GuSCN-based) | Inactivates virus, stabilizes RNA, and releases nucleic acids from crude samples (swabs, saliva). | Simplicity & Safety: Eliminates complex RNA extraction, reduces biohazard risk, and integrates with direct "sample-in, answer-out" workflows. |
| Portable Dry Bath / Heater | Provides precise isothermal incubation (65°C). | Infrastructure & Cost: Low-cost, battery-operated versions exist. Non-instrumented options (chemical heaters) are under research. |
| Synthetic SARS-CoV-2 RNA Control | Positive control for assay validation and monitoring. | Quality Control: Essential for establishing Limit of Detection (LoD) and verifying assay function in field conditions. |
Navigating Regulatory Approval (EUA, CE-IVD) for Diagnostic Deployment
This Application Note details the critical regulatory pathways for deploying diagnostic tests, specifically within the context of a thesis on the development of a Loop-Mediated Isothermal Amplification (LAMP) assay for SARS-CoV-2 detection. For researchers and developers, navigating Emergency Use Authorization (EUA) from the US FDA and Conformité Européenne In Vitro Diagnostic (CE-IVD) marking is essential for rapid clinical deployment during a public health emergency.
Regulatory Pathways: Comparison and Key Data
Table 1: Comparison of EUA and CE-IVD Pathways for SARS-CoV-2 Diagnostic Tests
| Aspect | FDA Emergency Use Authorization (EUA) | CE-IVD Marking (EU) |
|---|---|---|
| Legal Basis | Section 564 of the FD&C Act; Declared Public Health Emergency | In Vitro Diagnostic Regulation (IVDR) 2017/746 |
| Validity Period | Duration of the declared public health emergency | No expiry; continuous compliance required |
| Core Requirement | Demonstrated benefits outweigh known/potential risks | Conformity with Essential Safety & Performance Requirements |
| Performance Standards | Comparison to an authorized molecular assay (e.g., RT-PCR) | Fulfillment of Common Specifications (CS) for SARS-CoV-2 |
| Typical Review Timeline | ~14-60 days (expedited during pandemic) | Varies by Notified Body; several months |
| Key Clinical Study Metrics | Positive Percent Agreement (PPA) > 90-95%; Negative Percent Agreement (NPA) > 98-99% | Sensitivity > 90-95%; Specificity > 98-99% (per CS) |
| Quality System | Compliance with 21 CFR Part 820 (QSR) or ISO 13485 | ISO 13485 mandatory under IVDR |
| Post-Market Surveillance | Mandatory adverse event reporting (FAERS) | Vigilance system & Post-Market Performance Follow-up (PMPF) |
Table 2: Example Performance Data Required for a SARS-CoV-2 LAMP EUA Submission
| Study Parameter | Minimum Target (FDA Guideline) | Example LAMP Assay Results |
|---|---|---|
| Limit of Detection (LoD) | ≤ 10^4 copies/mL (or equivalent) | 5 RNA copies/µL (95% hit rate) |
| Clinical Sensitivity (PPA) | ≥ 90% (for symptomatic) | 94.1% (95% CI: 88.5%-97.0%) |
| Clinical Specificity (NPA) | ≥ 99% | 100% (95% CI: 96.8%-100%) |
| Inclusivity (Genetic Variants) | All known circulating strains | Tested against Alpha, Delta, Omicron (BA.1, BA.2) |
| Cross-Reactivity | No interference from common pathogens | Tested against 30 respiratory flora/viruses; no cross-reactivity |
| Sample Types | At least one anterior nasal swab | Nasal, Nasopharyngeal, Saliva validated |
Detailed Experimental Protocols
Protocol 1: Determination of Limit of Detection (LoD) for SARS-CoV-2 LAMP Assay
Objective: To establish the lowest concentration of SARS-CoV-2 RNA that can be reliably detected by the LAMP assay ≥95% of the time.
Materials (Research Reagent Solutions Toolkit):
- Synthetic SARS-CoV-2 RNA: Quantified genomic transcript (e.g., from Twist Bioscience) spanning N and E genes.
- LAMP Master Mix: Contains Bst 2.0/3.0 DNA polymerase, dNTPs, buffer, and stabilizers.
- Primer Set: Six primers (F3, B3, FIP, BIP, LF, LB) targeting conserved region of SARS-CoV-2 genome.
- Fluorescent Intercalating Dye: SYTO 9 or Calcein/MnCl2 for real-time or visual detection.
- Negative Matrix: Pooled human nasal swab transport media confirmed negative for SARS-CoV-2.
- Real-time Isothermal Fluorometer or Water Bath/Heating Block: Maintained at 65°C ± 1°C.
Procedure:
- Prepare a 10-fold serial dilution series of the SARS-CoV-2 RNA standard in negative matrix, ranging from 10^5 to 10^0 copies/µL.
- For each dilution level, prepare 20 replicates of the LAMP reaction.
- Reaction Volume: 25 µL.
- Components: 12.5 µL 2x Master Mix, 1 µL primer mix (final conc: 1.6 µM FIP/BIP, 0.2 µM F3/B3, 0.8 µM LF/LB), 5 µL RNA template, nuclease-free water to volume.
- Run reactions at 65°C for 30-40 minutes, monitoring fluorescence every 30-60 seconds or observing color change at endpoint.
- A positive result is defined by a fluorescence threshold time (Tt) < 30 min or a clear color change from orange to green/yellow.
- Calculate the hit rate for each dilution. The LoD is the lowest concentration where ≥19/20 (95%) replicates test positive.
Protocol 2: Clinical Agreement Study (vs. Authorized RT-PCR)
Objective: To determine Positive Percent Agreement (PPA) and Negative Percent Agreement (NPA) of the LAMP assay using residual patient specimens.
Materials:
- Clinical Specimens: De-identified anterior nasal swab specimens in transport media, stored at -80°C. Include at least 30 positive and 30 negative samples as determined by a reference EUA RT-PCR assay.
- Reference Method: An FDA-authorized SARS-CoV-2 RT-PCR assay (e.g., CDC 2019-nCoV RT-PCR).
- RNA Extraction Kit: (If required for LAMP protocol) or appropriate viral inactivation buffer for direct testing.
- LAMP Assay Components: As described in Protocol 1.
Procedure:
- Obtain IRB approval or waiver for use of residual, de-identified specimens.
- Aliquot specimens. Test all samples in a blinded manner with both the candidate LAMP assay and the reference RT-PCR assay.
- For the LAMP assay, follow the optimized procedure (e.g., direct addition of 5 µL of inactivated sample to the reaction mix from Protocol 1).
- Record all results. Resolve any discrepancies by retesting with both methods and/or sequencing.
- Construct a 2x2 concordance table. Calculate:
- PPA = [True Positives / (True Positives + False Negatives)] x 100
- NPA = [True Negatives / (True Negatives + False Positives)] x 100
Regulatory Workflow and Requirements Visualization
Diagram Title: EUA and CE-IVD Regulatory Workflow for Diagnostics
Diagram Title: Typical SARS-CoV-2 LAMP Testing Workflow
The Scientist's Toolkit: Essential Reagents & Materials
| Item | Function in SARS-CoV-2 LAMP Assay |
|---|---|
| Bst 2.0 or 3.0 DNA Polymerase | Thermostable enzyme for isothermal DNA amplification; lacks 5'→3' exonuclease activity. |
| LAMP Primer Set (F3, B3, FIP, BIP, LF, LB) | Six primers targeting 8 distinct regions of the SARS-CoV-2 genome for high specificity and rapid amplification. |
| Isothermal Amplification Buffer | Provides optimal pH, salt (MgSO4, (NH4)2SO4), and betaine conditions for Bst polymerase and strand displacement. |
| SYTO 9 or SYBR Green I Dye | Fluorescent intercalating dye for real-time monitoring of amplification on a fluorometer. |
| Calcein/MnCl2 with dNTPs | Colorimetric detection system; amplification depletes dNTPs, releasing Mn2+ to form bright green Calcein complex. |
| Synthetic SARS-CoV-2 RNA Control | Quantified positive control for LoD studies, assay validation, and routine quality control. |
| Human Specimen Matrix (Negative) | Validated negative nasal swab transport media for diluting standards and controlling for sample matrix effects. |
| Heat Block or Water Bath | Simple, low-cost device to maintain constant 60-65°C temperature required for LAMP reaction. |
Conclusion
The LAMP protocol for SARS-CoV-2 represents a powerful, adaptable tool that bridges the gap between complex laboratory PCR and rapid point-of-care testing. By understanding its foundational principles, meticulously following and optimizing the methodological protocol, proactively troubleshooting issues, and rigorously validating performance against standard benchmarks, researchers can develop robust assays. The future of LAMP in biomedical research extends beyond COVID-19, offering a versatile platform for detecting other pathogens and genetic markers in resource-limited settings. Continued innovation in primer design, lyophilization, and multiplexing will further solidify its role in pandemic preparedness and decentralized diagnostics. For scientists and drug developers, mastering LAMP technology is an investment in a flexible, rapid-response diagnostic capability with significant implications for global public health.