Decoding the SARS-CoV-2 Genome: A Comprehensive Guide to Diagnostic Protein Targets and Assay Development
This article provides a comprehensive technical overview of the SARS-CoV-2 genome structure and its encoded proteins, with a focused analysis on their application in molecular and antigen-based diagnostics.
Decoding the SARS-CoV-2 Genome: A Comprehensive Guide to Diagnostic Protein Targets and Assay Development
Abstract
This article provides a comprehensive technical overview of the SARS-CoV-2 genome structure and its encoded proteins, with a focused analysis on their application in molecular and antigen-based diagnostics. Targeting researchers, scientists, and drug development professionals, it details the genomic architecture from replicase complex to structural proteins, explores the principles and methodologies behind PCR, sequencing, and rapid antigen tests, addresses common challenges in assay design and optimization for emerging variants, and validates targets through comparative analysis of clinical performance and regulatory standards. The synthesis offers a strategic framework for developing robust diagnostic tools in a dynamically evolving viral landscape.
The SARS-CoV-2 Blueprint: Foundational Genome Architecture and Key Protein Functions
This whitepaper provides a technical overview of the SARS-CoV-2 genome, a ~30 kb positive-sense single-stranded RNA (+ssRNA) molecule. Framed within a thesis on viral genome structure and protein targets for diagnostics research, we detail the genomic organization, key functional elements, and experimental approaches for its study. This guide is intended for researchers, scientists, and drug development professionals engaged in virology and therapeutic discovery.
Genomic Organization and Key Features
The SARS-CoV-2 genome (NCBI Reference Sequence: NC_045512.2) is approximately 29.9 kilobases in length. It is 5’-capped and 3’-polyadenylated, enabling it to function directly as an mRNA for translation of the viral replicase polyproteins. The genome encodes both structural and non-structural proteins (nsps) in a defined order.
Primary Genomic Elements
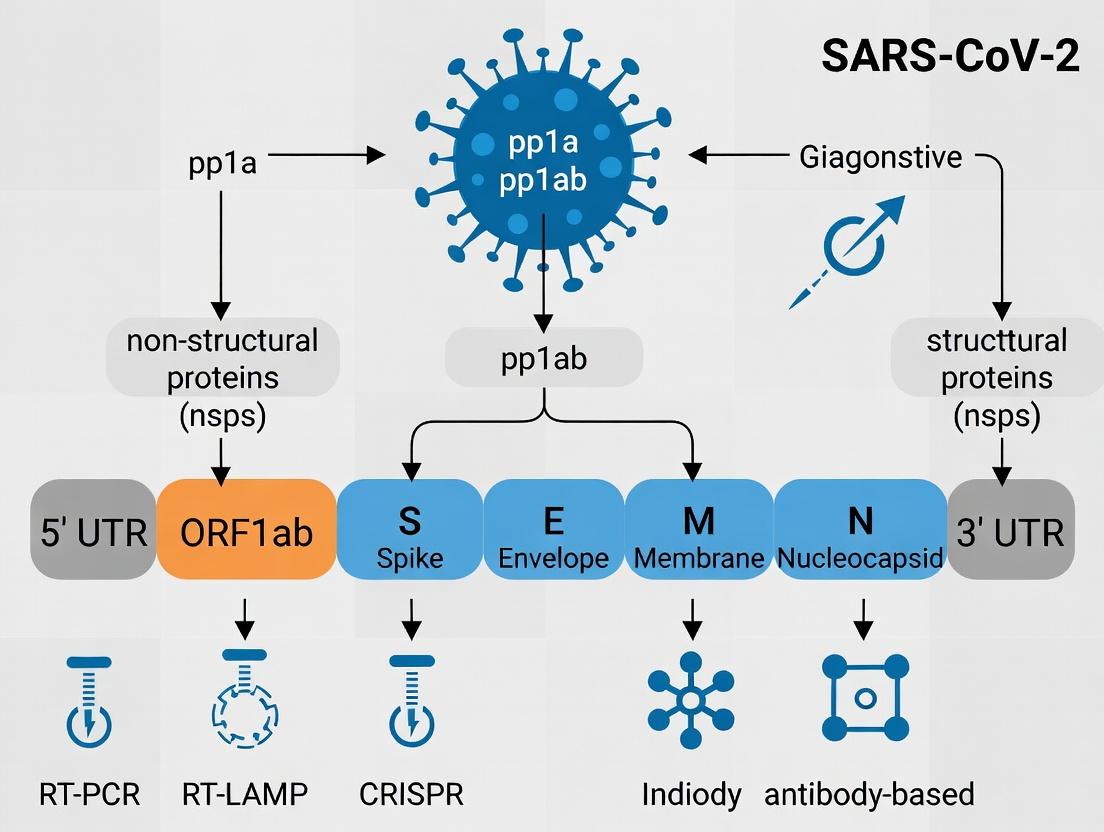
The genome contains 14 open reading frames (ORFs) flanked by untranslated regions (UTRs). Approximately two-thirds of the genome from the 5’ end comprises ORF1a and ORF1b, which are translated into polyproteins pp1a and pp1ab via a -1 ribosomal frameshift element (RFS). These are subsequently cleaved by viral proteases into 16 non-structural proteins (nsp1-16) that form the replication-transcription complex (RTC). The remaining one-third encodes the structural and accessory proteins.
Table 1: SARS-CoV-2 Genomic Regions and Functions
| Genomic Region | Nucleotide Position (approx.) | Encoded Product(s) | Primary Function(s) |
|---|---|---|---|
| 5’ UTR | 1-265 | N/A | Translation initiation, genome packaging |
| ORF1a | 266-13,483 | Polyprotein pp1a (nsp1-11) | Protease (nsp3, nsp5), RTC components |
| ORF1b (via RFS) | 13,468-21,563 | Polyprotein pp1ab (nsp1-16) | RNA-dependent RNA polymerase (RdRp, nsp12), Helicase (nsp13), Exonuclease (nsp14) |
| S Gene | 21,563-25,384 | Spike (S) glycoprotein | Host cell receptor binding, membrane fusion |
| ORF3a | 25,393-26,220 | ORF3a protein | Viroporin, modulation of host responses |
| E Gene | 26,245-26,472 | Envelope (E) protein | Virion assembly, budding, viroporin activity |
| M Gene | 26,523-27,191 | Membrane (M) protein | Central organizer of virion assembly |
| ORF6 | 27,202-27,387 | ORF6 protein | IFN antagonist, inhibits host nuclear import |
| ORF7a | 27,394-27,759 | ORF7a protein | Immune modulation, inhibits host protein synthesis |
| ORF7b | 27,756-27,887 | ORF7b protein | Unknown function |
| ORF8 | 27,894-28,259 | ORF8 protein | Immune evasion, modulates MHC-I |
| N Gene | 28,274-29,533 | Nucleocapsid (N) protein | RNA genome packaging, virion assembly, modulation of host cell processes |
| ORF10 | 29,558-29,674 | ORF10 (putative) | Function not fully characterized |
| 3’ UTR | 29,675-29,903 | N/A | Replication, transcription, polyadenylation |
Table 2: Key Quantitative Genomic Data
| Feature | Measurement | Notes / Significance |
|---|---|---|
| Genome Length | 29,903 nucleotides (nt) | Reference strain Wuhan-Hu-1 (NC_045512.2) |
| GC Content | ~38% | Lower than human genome (~41%), influences codon usage and secondary structure |
| ORF1a/b Length | 21,291 nt (pp1ab) | Encodes the replication-transcription complex |
| Frameshift Site Efficiency | ~40-70% | Regulates ratio of pp1a to pp1ab production |
| Subgenomic RNAs (sgRNAs) | 9-10 canonical species | Produced via discontinuous transcription; encode structural/accessory proteins |
| Mutation Rate | ~1 x 10⁻³ substitutions/site/year | Relatively low for an RNA virus due to proofreading by nsp14 (ExoN) |
Detailed Experimental Methodologies
Protocol for SARS-CoV-2 Genome Sequencing (Amplicon-Based Tiling PCR)
This protocol outlines the method for generating sequencing-ready libraries from viral RNA, commonly used for surveillance and variant tracking.
Principle: Reverse transcription followed by multiplex PCR using tiling primer pools to generate overlapping amplicons covering the entire viral genome.
Materials:
- Viral RNA extract (e.g., from nasopharyngeal swab)
- SuperScript IV Reverse Transcriptase (or equivalent)
- ARTIC Network version 4.1 primer pools (or latest version)
- Q5 Hot Start High-Fidelity DNA Polymerase
- AMPure XP beads
- Nextera XT or Illumina COVIDSeq library prep kit
- Illumina sequencing platform (MiSeq, NextSeq)
Procedure:
- Reverse Transcription: Combine 5.5 µL of viral RNA with 1 µL of random hexamers (50 ng/µL) and 1 µL of dNTPs (10 mM). Heat to 65°C for 5 min, then place on ice. Add 4 µL of 5X SSIV buffer, 1 µL of DTT (100 mM), 0.5 µL of RNaseOUT, and 1 µL of SuperScript IV. Incubate: 23°C for 10 min, 50°C for 10 min, 80°C for 10 min. Hold at 4°C.
- Multiplex PCR: Perform two separate PCR reactions using 2.5 µL of cDNA with Primer Pool 1 and Pool 2 (each pool contains ~100-125 primer pairs). Use Q5 polymerase with cycling: 98°C for 30 sec; 35 cycles of 98°C for 15 sec, 63°C for 5 min; final extension at 65°C for 5 min.
- PCR Clean-up: Pool the two PCR reactions and purify using 1X AMPure XP beads. Elute in 25 µL nuclease-free water.
- Library Preparation: Quantify purified amplicons by fluorometry. Use 5-10 ng as input for tagmentation and indexing with the Nextera XT kit, following manufacturer’s protocol.
- Sequencing: Pool indexed libraries, denature, and dilute to appropriate concentration for sequencing on an Illumina MiSeq (2x150 bp or 2x250 bp).
- Analysis: Demultiplex reads, trim adapters/primer sequences, perform de novo assembly or map to a reference genome (e.g., NC_045512.2) using pipelines like IVAR or Genome Detective.
Protocol for Pseudovirus Neutralization Assay
A key method for evaluating neutralizing antibodies against the SARS-CoV-2 Spike protein in diagnostic and vaccine sera.
Principle: A replication-incompetent viral vector (e.g., VSV-ΔG or lentivirus) is pseudotyped with the SARS-CoV-2 Spike protein. Neutralization of infectivity by serum antibodies is measured via a reporter gene (e.g., luciferase, GFP).
Materials:
- HEK293T-ACE2 cells (target cells expressing human ACE2 receptor)
- pCMV3-SARS2-S plasmid (expressing Wuhan-Hu-1 Spike, D614G, or variants)
- pNL4-3.Luc.R-E- lentiviral backbone (or VSV-ΔG-GFP/Fluc plasmid)
- Polyethylenimine (PEI) transfection reagent
- Test serum samples (heat-inactivated)
- Dulbecco’s Modified Eagle Medium (DMEM) with 10% FBS
- Bright-Glo Luciferase Assay System
Procedure:
- Pseudovirus Production:
- Seed HEK293T cells in a 10 cm dish. At 70-80% confluency, co-transfect with 10 µg of lentiviral backbone plasmid and 10 µg of Spike plasmid using PEI.
- 48-72 hours post-transfection, harvest supernatant containing pseudovirus particles. Centrifuge at 500 x g to remove cell debris, aliquot, and store at -80°C. Titer using HEK293T-ACE2 cells.
- Neutralization Assay:
- Serially dilute heat-inactivated serum samples (e.g., 4-fold dilutions starting at 1:20) in duplicate in a 96-well plate.
- Mix each serum dilution with an equal volume of pseudovirus (pre-titered to give ~100,000 RLU in assay). Incubate at 37°C for 1 hour.
- Add the serum-virus mixture to pre-seeded HEK293T-ACE2 cells in a 96-well plate.
- Incubate for 48-72 hours at 37°C, 5% CO₂.
- Detection:
- For luciferase reporter, aspirate medium, lyse cells, and add Bright-Glo substrate. Measure luminescence on a plate reader.
- Data Analysis: Calculate percent neutralization relative to virus-only control wells (0% neutralization) and cell-only wells (100% neutralization). Determine the serum dilution that inhibits 50% of infection (NT50 or IC50) using non-linear regression (e.g., four-parameter logistic curve) in GraphPad Prism.
Visualizations
Diagram 1: SARS-CoV-2 Genomic Expression and Replication Strategy (95 characters)
Diagram 2: Workflow for SARS-CoV-2 Whole Genome Sequencing (78 characters)
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Key Reagents for SARS-CoV-2 Genomic and Functional Research
| Reagent / Material | Vendor Examples | Primary Function in Research |
|---|---|---|
| SARS-CoV-2 Genomic RNA (Quantified) | BEI Resources, ATCC | Positive control for assay development (RT-qPCR, sequencing). |
| HEK293T-hACE2 Cell Line | InvivoGen, Sino Biological | Stable overexpression of human ACE2 receptor for viral entry, neutralization, and drug screening assays. |
| SARS-CoV-2 Spike Pseudotyping Kit | Integral Molecular, BPS Bioscience | For generating safe, replication-incompetent pseudoviruses to study viral entry and antibody neutralization. |
| Recombinant SARS-CoV-2 Proteins (S-RBD, N, 3CLpro) | ACROBiosystems, R&D Systems | Targets for ELISA development, antibody characterization, and high-throughput screening for inhibitors. |
| SARS-CoV-2 Genome qPCR Probe Assays | 2019-nCoV CDC Kit (IDT), Thermo Fisher TaqMan Assays | Quantitative detection of viral RNA (N1, N2, RdRp genes) for diagnostic research. |
| ARTIC Network Primers (V4.1+) | IDT, Swift Biosciences | Overlapping primer pools for robust, tiled amplification of the entire viral genome for sequencing. |
| Monoclonal Antibodies (Anti-S, Anti-N) | Cell Signaling Technology, GeneTex | Tools for Western blot, immunofluorescence, and ELISA to detect viral proteins in research samples. |
| Fluorescent RdRp/Nsp Inhibitors | Merck, MedChemExpress | Probe molecules for screening and characterizing inhibitors of the viral replication complex. |
| High-Fidelity Polymerase (Q5, Phusion) | NEB, Thermo Fisher | For accurate amplification of viral genomic segments with minimal error rates. |
| RNase Inhibitor (Murine, Recombinant) | NEB, Promega | Protection of viral RNA integrity during extraction and reverse transcription steps. |
Within the context of the SARS-CoV-2 genome structure and the identification of protein targets for diagnostics and therapeutic research, the replicase gene holds paramount importance. This gene, occupying approximately two-thirds of the viral RNA, is translated into two large polyproteins, pp1a and pp1ab, which are subsequently cleaved into 16 non-structural proteins (Nsps). These Nsps collectively form the replication-transcription complex (RTC), the core machinery responsible for viral RNA synthesis, proofreading, and subgenomic RNA production. This whitepaper provides an in-depth technical analysis of this machinery, its functions, and the experimental approaches used to study it.
Genomic Organization and Polyprotein Processing
The SARS-CoV-2 replicase gene consists of open reading frames 1a and 1b (ORF1a and ORF1b). A -1 ribosomal frameshift element between ORF1a and ORF1b controls the translation of the longer pp1ab polyprotein.
Table 1: SARS-CoV-2 Non-Structural Proteins (Nsps) and Primary Functions
| Nsp | Cleavage Sites (Approx.) | Key Function | Motifs/Domains |
|---|---|---|---|
| Nsp1 | pp1a 1-180 | Host translation inhibition, mRNA degradation | - |
| Nsp2 | pp1a 181-818 | Modulates host cell environment; precise function unclear | - |
| Nsp3 | pp1a 819-2763 | Papain-like protease (PLpro), deubiquitinase, macrodomain | Ubiquitin-like domain 1 (UB1), PLpro domain, Macrodomain (ADP-ribose binding) |
| Nsp4 | pp1a 2764-3263 | Induces membrane rearrangement; scaffold for RTC | Transmembrane domains |
| Nsp5 | pp1a 3264-3569 | Main protease (Mpro/3CLpro); cleaves viral polyproteins | Chymotrypsin-like protease domain |
| Nsp6 | pp1a 3570-3859 | Induces autophagosome-like vesicles; membrane curvature | Transmembrane domains |
| Nsp7 | pp1a 3860-3942 | Forms hexadecameric complex with Nsp8 as primase processivity factor | - |
| Nsp8 | pp1a 3943-4140 | Primase; forms hexadecameric complex with Nsp7 | - |
| Nsp9 | pp1a 4141-4253 | RNA-binding protein, dimerizes | Single-stranded RNA-binding |
| Nsp10 | pp1a 4254-4392 | Cofactor for Nsp14 and Nsp16; stimulates exoribonuclease and methyltransferase activities | - |
| Nsp11 | pp1a 4393-4405 | Short peptide; function unknown | - |
| Nsp12 | pp1ab 1-932 | RNA-dependent RNA polymerase (RdRp) | RdRp domain, NiRAN domain |
| Nsp13 | pp1ab 933-1272 | Helicase (5’ to 3’), RNA 5’ triphosphatase | NTPase/helicase domain, Zinc-binding domain |
| Nsp14 | pp1ab 1273-1584 | 3’->5’ exoribonuclease (ExoN) for proofreading, N7-guanine methyltransferase | ExoN domain, N7-MTase domain |
| Nsp15 | pp1ab 1585-1875 | Endoribonuclease (EndoU); evades host immune detection | Endouridylase domain |
| Nsp16 | pp1ab 1876-2096 | 2’-O-methyltransferase (2’-O-MTase); shields RNA from MDA5 recognition | 2’-O-MTase domain |
Processing is mediated by two viral proteases: Papain-like protease (PLpro) in Nsp3 cleaves at the Nsp1|2, Nsp2|3, and Nsp3|4 boundaries, while the main protease (Mpro, Nsp5) cleaves the remaining 11 sites. The mature Nsps then assemble into the RTC on double-membrane vesicles derived from the host endoplasmic reticulum.
Core Replication-Transcription Complex (RTC) Machinery and Functions
The Replication Core: Nsp12, Nsp7, Nsp8
The RNA-dependent RNA polymerase (RdRp, Nsp12) is the catalytic heart of the RTC. It requires Nsp7 and Nsp8 as cofactors for processive RNA synthesis. Nsp8 also exhibits primase activity.
Experimental Protocol 1: In Vitro RdRp Activity Assay
- Purpose: To measure the enzymatic activity of the purified Nsp12 (RdRp) complex.
- Reagents: Purified recombinant Nsp12, Nsp7, Nsp8; RNA template (e.g., poly(C)); nucleotide triphosphates (NTPs) including radiolabeled or fluorescently labeled GTP; reaction buffer (typically containing Mg2+, DTT, NaCl).
- Method:
- Assemble the core complex by incubating Nsp12, Nsp7, and Nsp8 in equimolar ratios.
- Set up a 50 µL reaction mixture containing reaction buffer, core complex, RNA template, and NTPs.
- Incubate at 30-37°C for 60 minutes.
- Stop the reaction by adding EDTA.
- Quantify synthesized RNA: For radioactive assays, use trichloroacetic acid precipitation and scintillation counting. For fluorescent assays, measure fluorescence after separating products via gel electrophoresis.
- Key Controls: Omission of template, NTPs, or RdRp complex; use of a known polymerase inhibitor (e.g., Remdesivir triphosphate).
The Proofreading Complex: Nsp14-Nsp10
Nsp14 provides 3’->5’ exoribonuclease (ExoN) activity critical for replication fidelity. Nsp10 acts as a cofactor, drastically stimulating ExoN activity.
The Capping Machinery: Nsp13, Nsp14, Nsp16-Nsp10
SARS-CoV-2 RNA is capped to mimic host mRNA. The pathway involves:
- RNA 5’ triphosphatase: Nsp13 hydrolyzes the 5’ γ-phosphate of nascent RNA.
- Guanylyltransferase: Likely a host enzyme adds GMP.
- N7-methyltransferase: The Nsp14 N7-MTase domain methylates the guanine cap.
- 2’-O-methyltransferase: The Nsp16-Nsp10 complex methylates the ribose 2’-O position, a step critical for immune evasion.
Diagram 1: Replicase Gene Processing and RTC Function (92 chars)
Key Experimental Methodologies for Studying Nsp Functions
Experimental Protocol 2: Co-Immunoprecipitation (Co-IP) for Nsp Complex Identification
- Purpose: To identify protein-protein interactions between Nsps (e.g., Nsp10 with Nsp14 or Nsp16).
- Reagents: Cells (e.g., HEK293T) transfected with tagged Nsp constructs (e.g., FLAG-Nsp10, HA-Nsp14); lysis buffer (non-denaturing, with protease inhibitors); anti-FLAG antibody-coupled beads; wash buffers; elution buffer (3X FLAG peptide or SDS sample buffer).
- Method:
- Co-transfect cells with plasmids expressing tagged Nsps.
- At 24-48 hours post-transfection, lyse cells in mild lysis buffer.
- Clarify lysate by centrifugation.
- Incubate supernatant with anti-FLAG magnetic beads for 2-4 hours at 4°C.
- Wash beads 3-5 times with wash buffer.
- Elute bound proteins using competitive FLAG peptide or by boiling in SDS sample buffer.
- Analyze eluates by western blot using anti-HA and anti-FLAG antibodies.
- Key Controls: Transfection with single tagged protein (for specificity), use of isotype control beads.
Experimental Protocol 3: Nsp16/Nsp10 2’-O-MTase Biochemical Assay
- Purpose: To quantify the methyltransferase activity of the Nsp16-Nsp10 complex.
- Reagents: Purified Nsp16-Nsp10 complex; short RNA substrate with a 5’ capped end (e.g., m7GpppA); S-adenosyl methionine (SAM) as methyl donor (can use 3H-labeled SAM for radioactivity); reaction buffer; scintillation fluid or separation columns.
- Method:
- In a reaction tube, mix Nsp16-Nsp10 complex, capped RNA substrate, and 3H-SAM in reaction buffer.
- Incubate at 30°C for 30-60 minutes.
- Stop the reaction (e.g., by heating or adding inhibitors).
- Separate the methylated RNA product from unincorporated 3H-SAM using spin columns or filter binding.
- Quantify radioactivity via scintillation counting to determine methyl group transfer.
- Key Controls: Omission of enzyme complex, RNA substrate, or SAM; use of a SAM-competitive inhibitor.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for SARS-CoV-2 Replicase/Nsp Research
| Reagent | Supplier Examples (Illustrative) | Function in Research |
|---|---|---|
| SARS-CoV-2 Nsp Expression Plasmids | Addgene, Sino Biological | Source for recombinant protein expression in bacterial, insect, or mammalian systems. |
| Recombinant SARS-CoV-2 Nsp Proteins | BPS Bioscience, RayBiotech, Abcam | For in vitro enzymatic assays (RdRp, protease, ExoN, MTase), binding studies, and inhibitor screening. |
| SARS-CoV-2 Replicon Cell Lines | Integral Molecular, Vero E6-based systems | Self-replicating viral RNA systems lacking structural genes; biosafe tools for studying replication and screening antivirals. |
| RdRp Inhibitors (e.g., Remdesivir-TP) | MedChemExpress, Cayman Chemical | Positive controls for in vitro RdRp activity assays and mechanism-of-action studies. |
| Mpro/PLpro Inhibitors (e.g., GC376, GRL0617) | Selleckchem, TargetMol | Controls for protease activity assays and polyprotein cleavage studies. |
| Anti-Nsp Antibodies (Monoclonal/Polyclonal) | GeneTex, Cell Signaling Technology, Genetex | Detection of Nsp expression in infected cells (IF, IHC), western blot analysis, and Co-IP experiments. |
| Fluorescent or Radioactive NTPs/SAM | PerkinElmer, Hartmann Analytic | Substrates for sensitive detection of polymerase or methyltransferase activity in kinetic assays. |
| Capped RNA Substrates | Trilink BioTechnologies, ChemGenes | Defined substrates for studying the viral capping enzymes Nsp14 and Nsp16. |
The replicase gene and its Nsp products constitute a highly coordinated molecular machine essential for SARS-CoV-2 replication and pathogenesis. Each component, from the core RdRp to the proofreading and capping enzymes, represents a validated target for antiviral drug development. Understanding their intricate structure, function, and interactions—through the rigorous experimental methodologies outlined—is fundamental to advancing diagnostic tools (e.g., targeting conserved Nsp sequences) and therapeutic interventions within the broader research thesis on SARS-CoV-2.
The SARS-CoV-2 virion is encased in a host-derived lipid bilayer studded with structural proteins: Spike (S), Membrane (M), Envelope (E), and Nucleocapsid (N). These proteins orchestrate virion assembly, release, and pathogenesis. Within the thesis context of SARS-CoV-2 genome structure and protein targets for diagnostics and therapeutic research, understanding these proteins is paramount. The S protein mediates viral entry and is a primary target for vaccines and therapeutics. The N protein packages the viral RNA genome and is a major target for diagnostic assays. The M and E proteins are critical scaffolds for virion assembly and budding. This whitepaper provides an in-depth technical analysis of their roles, supported by current experimental data and methodologies.
Protein Functions & Quantitative Data
Table 1: Core Structural Proteins of SARS-CoV-2
| Protein | Gene Location (nt) | Amino Acids | Molecular Weight (kDa) | Key Functions | Pathogenesis Role |
|---|---|---|---|---|---|
| Spike (S) | 21563-25384 | 1273 | ~180 (monomer) | Host cell receptor binding, membrane fusion, virion attachment. | Primary determinant of tropism and transmissibility; major antigen. |
| Nucleocapsid (N) | 28274-29533 | 419 | ~46 | RNA genome packaging, RNA synthesis modulation, host cell interference. | Inhibits host IFN response; major diagnostic antigen. |
| Membrane (M) | 26523-27191 | 222 | ~25-30 | Virion budding scaffold, curvature induction, interacts with all other structural proteins. | Modulates host immune response; critical for assembly efficiency. |
| Envelope (E) | 26245-26472 | 75 | ~8-12 | Virion assembly and budding, ion channel (viroporin) activity. | Promotes inflammation and pathogenesis; facilitates virion release. |
Table 2: Key Protein-Protein Interactions in Virion Assembly
| Interaction | Experimental Method (Common) | Functional Outcome |
|---|---|---|
| M-N | Co-immunoprecipitation, FRET | Packages ribonucleoprotein complex into budding virion. |
| M-S | Pull-down assays, Cryo-ET | Incorporates S trimers into viral envelope. |
| M-E | Yeast two-hybrid, BRET | Facilitates envelope curvature and budding. |
| M-M | Cross-linking, Molecular Modeling | Forms homodimers/oligomers; core scaffold lattice. |
| N-RNA | EMSA, NMR | Forms helical ribonucleoprotein complex for protection and transcription. |
Detailed Experimental Protocols
Protocol 1: Co-immunoprecipitation (Co-IP) for M-N Protein Interaction
Objective: To validate the physical interaction between the Membrane (M) and Nucleocapsid (N) proteins. Reagents: HEK-293T cells, plasmids expressing SARS-CoV-2 M-FLAG and N-HA, anti-FLAG M2 affinity gel, lysis buffer (50 mM Tris-HCl pH 7.4, 150 mM NaCl, 1% Triton X-100, protease inhibitors), wash buffer, 3xFLAG peptide for elution. Procedure:
- Transfection: Co-transfect HEK-293T cells with M-FLAG and N-HA expression plasmids using a polyethylenimine (PEI) method.
- Lysis: At 48h post-transfection, lyse cells in ice-cold lysis buffer for 30 min. Centrifuge at 16,000 x g for 15 min at 4°C.
- Pre-clearance: Incubate supernatant with protein A/G beads for 1h at 4°C to reduce non-specific binding.
- Immunoprecipitation: Incubate pre-cleared lysate with anti-FLAG M2 affinity gel overnight at 4°C with gentle rotation.
- Washing: Wash beads 5x with ice-cold wash buffer (lysis buffer with 0.1% Triton X-100).
- Elution: Elute bound proteins using 150 µg/mL 3xFLAG peptide in TBS.
- Analysis: Detect eluted proteins by SDS-PAGE and Western blotting using anti-FLAG and anti-HA antibodies.
Protocol 2: Electrophoretic Mobility Shift Assay (EMSA) for N Protein-RNA Binding
Objective: To assess the binding affinity of the Nucleocapsid (N) protein to specific SARS-CoV-2 genomic RNA sequences. Reagents: Purified recombinant N protein, 5'-Cy5-labeled RNA probe (e.g., from the N gene packaging signal), unlabeled specific and nonspecific competitor RNA, binding buffer (10 mM HEPES pH 7.3, 50 mM KCl, 1 mM MgCl2, 0.5 mM DTT, 0.05% NP-40, 5% glycerol), 6% native polyacrylamide gel. Procedure:
- Probe Preparation: Anneal the labeled single-stranded RNA probe.
- Binding Reaction: Incubate increasing concentrations of N protein (0-2 µM) with a fixed amount of labeled RNA probe (10 nM) in binding buffer for 30 min at room temperature. Include reactions with excess unlabeled competitor to test specificity.
- Electrophoresis: Load reactions onto a pre-run 6% native PAGE gel in 0.5x TBE buffer. Run at 100V for 60-90 min at 4°C.
- Visualization: Image the gel using a fluorescence scanner for the Cy5 label. A shift in the RNA probe's mobility indicates protein-RNA complex formation.
Visualization: Pathways and Workflows
Diagram 1: SARS-CoV-2 Structural Protein Assembly & Budding Pathway (83 chars)
Diagram 2: Co-IP Workflow for M-N Protein Interaction (55 chars)
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Reagents for Structural Protein Studies
| Reagent/Material | Supplier Examples | Function in Research |
|---|---|---|
| HEK-293T Cells | ATCC, Thermo Fisher | Highly transfectable mammalian cell line for protein expression and interaction studies. |
| pcDNA3.1(+) Expression Vector | Invitrogen, Addgene | Common backbone for cloning and expressing SARS-CoV-2 structural protein genes. |
| Anti-SARS-CoV-2 S (RBD) mAb | Sino Biological, GeneTex | Neutralizing antibody for S protein function-blocking assays and detection. |
| Recombinant N Protein (full-length) | Abcam, RayBiotech | Positive control for ELISA, EMSA, and serology assay development. |
| Anti-FLAG M2 Affinity Gel | Sigma-Aldrich | For immunoprecipitation of FLAG-tagged proteins (e.g., M-FLAG). |
| Lipofectamine 3000 | Thermo Fisher | High-efficiency transfection reagent for plasmid delivery into cells. |
| Protease Inhibitor Cocktail | Roche, Thermo Fisher | Prevents protein degradation during cell lysis and protein purification. |
| SuperScript IV RT | Thermo Fisher | Reverse transcriptase for generating cDNA from viral RNA for cloning. |
| NativePAGE Bis-Tris Gels | Thermo Fisher | For running EMSAs and analyzing native protein-RNA complexes. |
| Cryo-Electron Microscopy Grids | Quantifoil, Thermo Fisher | Support film for visualizing virion structure and protein complexes at near-atomic resolution. |
Within the context of SARS-CoV-2 research, accessory proteins (ORF3a, ORF6, ORF7a, ORF7b, ORF8, ORF9b, ORF10) are non-structural elements critical for viral pathogenesis. This whitepaper details their mechanisms in immune evasion, quantitative impacts on host pathways, and their emerging potential as targets for novel diagnostic platforms. Framed within a thesis on viral genome structure, we position these proteins as key to understanding infectivity and developing next-generation detection assays.
Mechanism of Action: Immune Modulation by SARS-CoV-2 Accessory Proteins
Accessory proteins function as versatile modulators, targeting multiple nodes of innate immune signaling, particularly the type I interferon (IFN-I) response.
Diagram: SARS-CoV-2 Accessory Proteins Inhibit IFN-I Signaling Cascade
Table 1: Quantitative Impact of Key SARS-CoV-2 Accessory Proteins on Immune Markers
| Accessory Protein | Primary Immune Target | Reported Effect Size (In Vitro) | Experimental System |
|---|---|---|---|
| ORF6 | Nuclear import (STAT1, IRF3) | >80% reduction in IFN-β mRNA | HEK-293T, A549 cells |
| ORF9b | MAVS, RIG-I, TOM70 | ~70% suppression of IFN-β promoter activity | HEK-293T, PBMCs |
| ORF3a | NLRP3 Inflammasome | 2-3 fold increase in IL-1β secretion | THP-1 macrophages |
| ORF8 | MHC-I degradation | ~50% reduction in surface MHC-I | HEK-293T, A549 cells |
| ORF7a | Blocks STAT2 phosphorylation | ~60% reduction in ISRE promoter activity | HeLa cells |
Experimental Protocols for Characterizing Accessory Protein Function
Protocol: Luciferase Reporter Assay for IFN Pathway Inhibition
Objective: Quantify the inhibitory effect of an accessory protein on IFN-β promoter activity.
- Cell Seeding: Seed HEK-293T cells in a 24-well plate at 1x10^5 cells/well and incubate for 24h.
- Transfection: Co-transfect using a polyethylenimine (PEI) protocol:
- Test Plasmid: 200 ng pcDNA3.1-ORF (e.g., ORF6, ORF9b).
- Reporter Plasmid: 100 ng pGL4-IFN-β-promoter-firefly-luciferase.
- Induction Plasmid: 50 ng pUNO-hRIG-I (constitutively active).
- Control Plasmid: 20 ng pRL-TK-Renilla-luciferase for normalization.
- Incubation: Harvest cells 48 hours post-transfection.
- Lysis & Measurement: Lyse cells with Passive Lysis Buffer (Promega). Measure Firefly and Renilla luciferase activity using a dual-luciferase reporter assay system on a microplate luminometer.
- Analysis: Normalize Firefly luminescence to Renilla luminescence. Express data as fold-change relative to cells transfected with empty vector control.
Protocol: Flow Cytometry for Surface MHC-I Downregulation
Objective: Assess ORF8-mediated degradation of MHC-I molecules.
- Transfection: Transfect A549 cells with a plasmid expressing ORF8-FLAG or empty vector.
- Incubation: Incubate for 36-48 hours.
- Staining: Harvest cells, wash with PBS+2% FBS. Stain with APC-conjugated anti-HLA-A,B,C antibody (or isotype control) for 30 min on ice in the dark.
- Fixation: Wash cells and fix with 2% paraformaldehyde.
- Acquisition: Analyze on a flow cytometer. Gate on live, single cells. Compare the median fluorescence intensity (MFI) of the ORF8-expressing population versus the vector control.
- Validation: Parallel wells can be lysed for Western blot to confirm ORF8 expression and correlate with MHC-I protein levels.
Diagram: Experimental Workflow for Immune Modulation Assays
Diagnostic Relevance and Research Toolkit
Accessory proteins, due to their critical roles and unique sequences, offer targets for differentiating SARS-CoV-2 from other coronaviruses and detecting active infection.
Table 2: Accessory Proteins as Diagnostic Targets
| Target Protein | Diagnostic Modality | Advantage | Challenge |
|---|---|---|---|
| ORF8 | Antigen-detecting lateral flow assay (LFA) | Early expression, elicits strong antibody response | Sequence variability (deletions observed) |
| ORF3a | Electrochemical immunosensor | High surface expression in infected cells | Cross-reactivity with other betacoronavirus proteins? |
| ORF6/ORF9b | RT-PCR multiplex assay | High sequence specificity for SARS-CoV-2 | Low abundance RNA requires sensitive amplification |
The Scientist's Toolkit: Key Research Reagents
| Reagent/Material | Provider Examples | Function in This Field |
|---|---|---|
| pcDNA3.1-SARS2-ORF Plasmids | Addgene, Sino Biological | Mammalian expression of accessory proteins for functional studies. |
| Dual-Luciferase Reporter Assay System | Promega | Quantifies promoter activity (e.g., IFN-β) for immune modulation assays. |
| Anti-FLAG M2 Magnetic Beads | Sigma-Aldrich | Immunoprecipitation of tagged accessory proteins for interactome studies. |
| Human IFN-β ELISA Kit | PBL Assay Science | Measures secreted IFN-β to validate pathway inhibition. |
| APC anti-human HLA-A,B,C Antibody | BioLegend | Flow cytometry staining to quantify MHC-I surface expression. |
| SARS-CoV-2 (ORF8) Monoclonal Antibody | Invitrogen, GeneTex | Key reagent for developing ORF8-specific antigen capture assays. |
| NLRP3 Inhibitor (MCC950) | Cayman Chemical | Tool compound to validate ORF3a's inflammasome activation mechanism. |
Within the SARS-CoV-2 genome, the distribution of genetic variation is non-uniform. "Genomic hotspots" refer to loci with a propensity for mutation, often within key structural or functional protein domains. Conversely, "conserved regions" remain invariant across lineages and are critical for viral viability. This whitepaper, framed within a thesis on SARS-CoV-2 genome structure and protein targets for diagnostics, delineates these regions, analyzes the molecular impact of Variants of Concern (VoCs), and provides technical guidance for related research.
Genomic Landscape: Conserved Regions vs. Variable Domains
The ~30 kb positive-sense single-stranded RNA genome of SARS-CoV-2 encodes both structural and non-structural proteins. Conserved regions are typically essential for replication and basic viral function, while variable domains often involve host-interaction surfaces under selective immune pressure.
Table 1: Key Conserved Genomic Regions in SARS-CoV-2
| Genomic Region | ORF/Protein | Nucleotide Position (approx.) | Functional Role | Conservation Rationale |
|---|---|---|---|---|
| RdRp Active Site | nsp12 (RdRp) | 13,442-16,236 | Viral RNA replication | Catalytic core; mutations often deleterious. |
| Proofreading Exonuclease | nsp14 | 18,075-19,620 | RNA proofreading & fidelity | Essential for replication complex integrity. |
| 5'-UTR & 3'-UTR | N/A | 1-265 & 29,674-29,903 | Genome replication/translation initiation | Structured RNA elements for ribosomal entry and replication. |
| Main Protease Active Site | nsp5 (Mpro/3CLpro) | 9,860-10,954 | Polyprotein processing | Precise catalytic triad required for cleavage. |
Table 2: Primary Variable Domains & Hotspots in SARS-CoV-2
| Genomic Region | ORF/Protein | Nucleotide Position (approx.) | Variable Domain/Hotspot | Functional Consequence |
|---|---|---|---|---|
| Spike Glycoprotein | S | 21,563-25,384 | Receptor-Binding Domain (RBD), N-Terminal Domain (NTD) | Alters ACE2 affinity, antibody neutralization. |
| Nucleocapsid Protein | N | 28,274-29,533 | Linker Region (LKR) | Affects RNA binding, protein oligomerization, and immunodominance. |
| Spike Glycoprotein | S | 21,563-25,384 | Furin Cleavage Site (FCS) | Impacts spike priming and infectivity. |
| ORF8 | ORF8 | 27,894-28,259 | Dimerization interface | Modulates host immune response (e.g., interferon antagonism). |
Impact of Variants of Concern (VoCs)
VoCs are defined by mutations in hotspots that confer a selective advantage, such as increased transmissibility, immune evasion, or disease severity. The following table summarizes key VoC-defining mutations in genomic hotspots.
Table 3: Defining Mutations in Selected SARS-CoV-2 VoCs (2022-2024)
| VoC (Pango Lineage) | Key Spike Mutations in Hotspots | Key Non-Spike Mutations | Impact on Phenotype |
|---|---|---|---|
| Omicron BA.5 (B.1.1.529.5) | G339D, S371F, S373P, S375F, T376A, D405N, R408S, K417N, N440K, L452R, S477N, T478K, E484A, F486V, Q498R, N501Y, Y505H | N:P151S, ORF6:D61L, nsp4:L438F, nsp5:P132H | Significant immune escape, increased ACE2 affinity. |
| Omicron XBB.1.5 (XBB.1.5) | R346T, K417N, V445P, G446S, N460K, F486P, F490S | N:T24I, ORF9b:T40A, nsp2:K81N | Enhanced receptor binding, evasion of neutralizing antibodies. |
| Omicron JN.1 (BA.2.86.1.1) | R346T, S371L, W152R, F157L, L455S | ORF1a:V621F, ORF1b:D277Y | Increased immune evasion and transmissibility. |
| Delta (B.1.617.2) | L452R, T478K, P681R | N:D63G, N:R203M, N:D377Y | Increased fusogenicity, viral load, and severity. |
Experimental Protocols for Analysis
Protocol: Deep Sequencing and Variant Calling for Hotspot Identification
Objective: To identify low-frequency variants and map genomic hotspots from clinical samples. Methodology:
- RNA Extraction & Library Prep: Use high-fidelity RT-PCR (e.g., ARTIC network v4.1 primer pools) for tiling amplicon generation. Convert to sequencing library using ligation-based kits (e.g., Illumina DNA Prep).
- Sequencing: Perform paired-end sequencing (2x150 bp) on an Illumina MiSeq or NextSeq platform to achieve >1000x depth.
- Bioinformatic Analysis:
- Quality Control & Trimming: Use FastQC and Trimmomatic.
- Alignment: Map reads to reference genome (NC_045512.2) using BWA-MEM.
- Variant Calling: Identify SNPs and indels using iVar (min-quality 20, min-frequency 0.05). Call consensus sequence.
- Hotspot Analysis: Use Nextclade for lineage assignment and SnpEff for functional annotation of variants. Visualize variant frequency across the genome with Python (Matplotlib).
Protocol: Pseudovirus Neutralization Assay for VoC Impact
Objective: Quantify the neutralizing antibody escape of VoC-specific spike mutations. Methodology:
- Pseudovirus Production: Co-transfect HEK293T cells with a lentiviral backbone (e.g., pNL4-3.Luc.R-E-) and a plasmid expressing the VoC spike protein using polyethylenimine (PEI).
- Harvesting: Collect supernatant at 48-72 hours post-transfection, filter (0.45 µm), aliquot, and store at -80°C.
- Neutralization Assay:
- Serially dilute test serum or monoclonal antibodies in a 96-well plate.
- Incubate with a standardized pseudovirus dose (MOI ~0.1) for 1 hour at 37°C.
- Add HEK293T-ACE2 cells and incubate for 48-72 hours.
- Lyse cells and measure luciferase activity.
- Analysis: Calculate the 50% neutralization titer (NT50) or inhibitory concentration (IC50) using a 4-parameter logistic curve in GraphPad Prism.
Diagram Title: Workflow for Genomic Hotspot Identification
Diagram Title: Pseudovirus Neutralization Assay Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Research Reagents for VoC and Hotspot Analysis
| Reagent/Category | Example Product (Supplier) | Function in Research |
|---|---|---|
| High-Fidelity PCR Mix | LunaScript RT SuperMix (NEB) or Q5 Hot Start (NEB) | Accurate amplification of viral RNA/cDNA for sequencing and cloning. |
| ARTIC Primers | ARTIC nCoV-2019 V4.1 Panel (IDT) | Tiling amplicon generation for comprehensive genome sequencing. |
| Spike Expression Plasmids | VoC Spike pcDNA3.1 vectors (BEI Resources) | Source of mutant spike for pseudovirus or protein production. |
| Lentiviral Backbone | pNL4-3.Luc.R-E- (NIH AIDS Reagent Program) | Reporter pseudovirus backbone for neutralization assays. |
| ACE2-Expressing Cell Line | HEK293T-hACE2 (BEI Resources or commercial) | Target cell line for pseudovirus entry and neutralization assays. |
| SARS-CoV-2 Nucleocapsid Antibody | Anti-SARS-CoV-2 Nucleocapsid mAb (Sino Biological) | Detection of viral protein in western blot or immunofluorescence. |
| Human Convalescent Serum Panel | WHO International Standard (NIBSC) | Reference standard for neutralizing antibody assays. |
| Bioinformatic Pipeline | iVar, Nextclade, SnpEff (Open Source) | Critical software for variant calling, lineage assignment, and annotation. |
From Sequence to Test: Methodologies for Targeting SARS-CoV-2 Proteins in Diagnostic Assays
The SARS-CoV-2 genome is a positive-sense, single-stranded RNA of approximately 30 kb. Effective diagnostic assays rely on precise targeting of conserved, essential genomic regions. This whitepaper details the design of primers and probes for Reverse Transcription Quantitative Polymerase Chain Reaction (RT-qPCR) targeting the RNA-dependent RNA polymerase (RdRp), Nucleocapsid (N), Envelope (E), and Spike (S) genes. These targets are central to a broader thesis on SARS-CoV-2 genome structure, function, and the strategic selection of protein-coding sequences for diagnostic and therapeutic research.
Target Gene Selection Rationale
| Gene Target | Genomic Function | Rationale for Diagnostic Targeting | Conservation Level |
|---|---|---|---|
| RdRp (ORF1ab) | Viral replication enzyme | Highly conserved; critical for virus viability, minimizing escape mutations. | Very High |
| N | Packages viral RNA | Highly expressed; abundant RNA transcript, increasing assay sensitivity. | High |
| E | Envelope protein formation | Small, highly conserved gene; excellent screening target. | Very High |
| S | Host cell receptor binding | Important for variant tracking; less conserved than others. | Moderate |
Primer & Probe Design Parameters
Key design principles ensure specificity, sensitivity, and robustness:
- Amplicon Length: 70-120 bp for optimal PCR efficiency.
- Tm: Primer Tm: 58-60°C; Probe Tm: 68-70°C (8-10°C higher than primers).
- Specificity: Verify against human and common respiratory pathogen genomes via BLAST.
- Variant Consideration: Design within conserved blocks, avoiding known variant sites (especially for S gene).
- Chemistry: Typically uses 5' fluorophore (e.g., FAM), 3' quencher (e.g., BHQ1), and a minor groove binder (MGB) or similar to enhance binding.
Experimentally Validated Primer/Probe Sequences
The following table summarizes key sequences from authoritative public health protocols (e.g., WHO, CDC).
Table 1: Recommended Primer and Probe Sets for SARS-CoV-2 RT-qPCR
| Target Gene | Assay Name (Source) | Sequence (5' -> 3') | Final Conc. (nM) | Amplicon (bp) |
|---|---|---|---|---|
| RdRp | Charité RdRp_SARSr-P1 (WHO) | F: GTGARATGGTCATGTGTGGCGGR: CARATGTTAAASACACTATTAGCATAP: FAM-CCAGGTGGWACRTCATCMGGTGATGC-BHQ1 | 400 (F/R)200 (P) | 155 |
| N | CDC N1 (USA) | F: GACCCCAAAATCAGCGAAATR: TCTGGTTACTGCCAGTTGAATCTGP: FAM-ACCCCGCATTACGTTTGGTGGACC-BHQ1 | 500 (F/R)250 (P) | 72 |
| E | Charité E (WHO) | F: ACAGGTACGTTAATAGTTAATAGCGTR: ATATTGCAGCAGTACGCACACAP: FAM-ACACTAGCCATCCTTACTGCGCTTCG-BHQ1 | 400 (F/R)200 (P) | 113 |
| S | (Example for Variants) | F: CTGGTGCAGTTCAAGTGTGTR: CAGACATTTTGCTCTCAAGCTGP: FAM-CACATTGGCCCCCAAATCCT-MGB | 400 (F/R)200 (P) | ~100 |
Experimental Protocol: One-Step RT-qPCR Assay
This protocol assumes extraction of viral RNA and use of a one-step RT-qPCR master mix.
I. Reagent Setup (25 µL Reaction)
| Component | Volume (µL) | Function |
|---|---|---|
| 2X One-Step RT-qPCR Master Mix | 12.5 | Contains polymerase, reverse transcriptase, dNTPs, buffer, Mg2+. |
| Forward Primer (20 µM) | 0.625 | Final conc: 500 nM. |
| Reverse Primer (20 µM) | 0.625 | Final conc: 500 nM. |
| Probe (10 µM) | 0.5 | Final conc: 200 nM. |
| RNase-free Water | 5.75 | Carrier. |
| Template RNA | 5 | Up to 5 µL of extracted RNA. |
| Total Volume | 25 |
II. Thermal Cycling Conditions (Standard)
| Step | Temperature | Time | Cycles | Purpose |
|---|---|---|---|---|
| Reverse Transcription | 50°C | 10-15 min | 1 | cDNA synthesis. |
| Initial Denaturation | 95°C | 2-3 min | 1 | Enzyme activation, template denaturation. |
| Amplification | 95°C | 10-15 sec | 40-45 | Denaturation. |
| 55-60°C | 30-60 sec | Annealing/Extension & Fluorescence Acquisition. | ||
| Hold | 4-10°C | ∞ |
III. Data Analysis
- Threshold: Set in the exponential phase of amplification.
- Ct (Cycle threshold): Positive results are typically Ct < 40.
- Controls: Include no-template control (NTC), positive control, and extraction control.
Visualizing Assay Workflow and Genomic Targets
Diagram 1: RT-qPCR Assay Workflow for SARS-CoV-2 Detection
Diagram 2: Genomic Targets for SARS-CoV-2 Diagnostic Primers
The Scientist's Toolkit: Essential Research Reagent Solutions
| Category | Item/Reagent | Function & Rationale |
|---|---|---|
| Core Enzymes | One-Step RT-qPCR Master Mix (e.g., TaqPath, Luna) | Integrated reverse transcriptase and hot-start DNA polymerase for streamlined, sensitive detection in closed-tube reactions. |
| Primers & Probes | HPLC- or PAGE-purified Oligonucleotides | High-purity primers and dual-labeled probes (FAM/BHQ1) minimize synthesis errors and background fluorescence. |
| RNA Handling | RNase Inhibitor | Protects viral RNA templates from degradation during setup. |
| Magnetic Bead-based RNA Extraction Kit (e.g., silica membrane) | Efficient, high-throughput purification of viral RNA from swab/transport media. | |
| Standards & Controls | Synthetic SARS-CoV-2 RNA Control (full-length or fragment) | Quantitative standard for generating standard curves (copy number determination) and positive run control. |
| Exogenous Internal Control (e.g., MS2 phage, human RNase P) | Co-extracted and co-amplified control to monitor extraction efficiency and rule out PCR inhibition. | |
| Consumables | Nuclease-free Tubes, Tips, and Plates | Prevents sample degradation and cross-contamination. |
| Software | Primer Design Software (e.g., Primer-BLAST, Geneious) | Identifies specific, efficient primer/probe sets with checks for secondary structure. |
| qPCR Analysis Software (instrument-specific) | For Ct determination, quantification, and quality assessment. |
This whitepaper details sequencing methodologies critical for a broader thesis on SARS-CoV-2 genome structure and protein targets for diagnostics research. Understanding viral evolution through variant surveillance is foundational to identifying conserved genomic regions and mutable protein domains (e.g., Spike RBD, Nucleocapsid) for diagnostic and therapeutic targeting.
Core Sequencing Technologies
Whole Genome Sequencing (WGS)
WGS provides a comprehensive, unbiased view of the entire viral genome (~29.9 kb for SARS-CoV-2). It is the gold standard for de novo variant discovery and characterizing complex genomic rearrangements.
Key Protocol (Illumina COVIDSeq Test):
- Reverse Transcription: Convert viral RNA to cDNA using random hexamers and reverse transcriptase.
- Multiplex PCR Amplification: Amplify the entire genome in overlapping amplicons using a primer panel (e.g., ARTIC Network v4.1 primer set).
- Library Preparation: Fragment amplicons, add Illumina sequencing adapters and dual-index barcodes via tagmentation (Nextera XT).
- Sequencing: Pool libraries and sequence on an Illumina platform (MiSeq, NextSeq) using 2x150 bp paired-end chemistry.
- Bioinformatics: Demultiplex reads, trim adapters, map to reference genome (MN908947.3), call variants, and perform phylogenetic analysis.
Targeted Amplicon Sequencing
This method uses PCR to enrich specific genomic regions prior to sequencing, offering high sensitivity and depth for variant detection in mixed samples, ideal for surveillance.
Key Protocol (ARTIC Network Workflow):
- Reverse Transcription: As above.
- Two-Pool PCR Amplification: Amplify the genome using two multiplexed primer pools (each covering tiled, overlapping ~400 bp amplicons) to minimize primer competition and amplicon dropout.
- Amplicon Purification: Clean up PCR products using magnetic beads (e.g., SPRIselect).
- Library Preparation & Sequencing: Quantify amplicons, proceed with library prep (often using ligation-based kits), and sequence on Illumina or Oxford Nanopore platforms.
Quantitative Data Comparison
Table 1: Technical Comparison of WGS and Amplicon Sequencing for SARS-CoV-2 Surveillance
| Parameter | Whole Genome Sequencing (Shotgun) | Targeted Amplicon Sequencing |
|---|---|---|
| Primary Use Case | De novo discovery, complex variants, metagenomics | High-throughput variant monitoring, low viral load samples |
| Input RNA | 10 - 1000 copies/µL | 1 - 100 copies/µL (higher effective sensitivity) |
| Typical Coverage Depth | 100x - 1000x | 1000x - 100,000x |
| Key Advantage | Unbiased; detects unknowns | Highly sensitive and robust; lower cost per sample |
| Key Limitation | Lower sensitivity in mixed samples; host contamination | Primer mismatches can cause amplicon failure; limited to targeted region |
| Time to Result | ~24-48 hours | ~12-24 hours |
| Cost per Sample (approx.) | $50 - $150 | $20 - $80 |
Table 2: Common SARS-CoV-2 Sequencing Platforms (2024)
| Platform | Technology | Read Length | Output per Run | Best Suited For |
|---|---|---|---|---|
| Illumina NextSeq 2000 | Sequencing-by-Synthesis | Up to 2x150 bp | 100-400 Gb | High-throughput population surveillance |
| Oxford Nanopore MinION | Nanopore Sensing | Ultra-long (>10 kb) | 10-50 Gb | Rapid, real-time surveillance in field labs |
| Illumina MiSeq | Sequencing-by-Synthesis | Up to 2x300 bp | 0.3-15 Gb | Small-batch validation & development |
| Ion Torrent Genexus | Semiconductor pH | Up to 400 bp | 1-15 Gb | Automated, hands-off WGS |
Visualization of Workflows
WGS for SARS-CoV-2 Workflow (Shotgun)
Amplicon Sequencing Workflow for SARS-CoV-2
Sequencing Method Decision Logic
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for SARS-CoV-2 Sequencing
| Item | Example Product | Function in Workflow |
|---|---|---|
| Viral RNA Extraction Kit | QIAamp Viral RNA Mini Kit, MagMAX Viral/Pathogen Kit | Isolates high-quality viral RNA from clinical samples (swab, saliva). |
| Reverse Transcriptase | SuperScript IV, LunaScript RT | Converts labile viral RNA to stable cDNA for downstream amplification. |
| High-Fidelity DNA Polymerase | Q5 Hot Start, PrimeSTAR GXL | Ensures accurate amplification of viral genome with minimal errors. |
| Tiled Primer Panels | ARTIC Network V4.1, Midnight Panel | Pre-designed primer sets for multiplex PCR covering entire SARS-CoV-2 genome. |
| Library Prep Kit | Illumina COVIDSeq Test, Nextera XT | Prepares amplicons for sequencing by adding adapters and indexes. |
| Sequence Capture Beads | SPRIselect, AMPure XP | Size-selects and purifies DNA fragments (amplicons, libraries). |
| Sequencing Control | SARS-CoV-2 RNA Genome Control | Positive control to validate entire workflow from extraction to sequencing. |
| Bioinformatics Pipeline | nf-core/viralrecon, ARTIC bioinformatics | Automated pipeline for read QC, variant calling, and lineage reporting. |
Within the broader thesis on SARS-CoV-2 genome structure and diagnostic targets, this document focuses on the translational application of structural knowledge. The SARS-CoV-2 genome encodes four major structural proteins: Spike (S), Envelope (E), Membrane (M), and Nucleocapsid (N). For antigen-based diagnostics, the N and S proteins are the primary targets. The N protein, abundant and highly conserved, is ideal for capturing viral presence. The S protein, specifically its receptor-binding domain (RBD), is critical for detecting infectious potential and vaccine-induced immune responses. This whitepaper details the immunoassay principles leveraging these targets.
Core Principles of Immunoassays and Lateral Flow Technology
Immunoassay Foundation
Immunoassays are biochemical tests that measure the presence or concentration of an analyte (antigen) through antigen-antibody (Ag-Ab) reactions. The fundamental components include:
- Capture Antibody: Immobilized on a solid surface.
- Detection Antibody: Conjugated to a label (e.g., gold nanoparticles, latex beads, enzymes).
- Antigen: The target molecule (e.g., SARS-CoV-2 N or S protein).
Lateral Flow Assay (LFA) Architecture
LFAs are a subset of immunoassays designed for rapid, point-of-care use. Key components are arranged on a nitrocellulose strip:
- Sample Pad: Where the sample is applied.
- Conjugate Pad: Contains labeled detection antibodies (e.g., colloidal gold-anti-N antibody) that bind the target antigen.
- Nitrocellulose Membrane: Contains immobilized capture lines.
- Test Line (T): Coated with a second, capture antibody specific to the target antigen (e.g., another anti-N antibody).
- Control Line (C): Coated with antibodies that capture the labeled detection antibody irrespective of antigen presence.
- Absorbent Pad: Acts as a sink to draw fluid through the strip via capillary action.
Diagram 1: LFA Process Flow (96 chars)
Quantitative Comparison of N and S Protein Targets
The selection of N or S protein as the target antigen involves trade-offs in sensitivity, specificity, and utility.
Table 1: Comparative Analysis of SARS-CoV-2 Antigen Targets for Immunoassays
| Parameter | Nucleocapsid (N) Protein | Spike (S) Protein / RBD |
|---|---|---|
| Abundance in Virion | High (~1000 copies/virion) | Moderate (~25-40 trimers/virion) |
| Conservation | High (slower mutation rate) | Lower (subject to selective pressure, variants) |
| Primary Diagnostic Utility | Detection of active infection | Detection of infection; surrogate for neutralizing antibodies |
| Typical LFA Limit of Detection (LoD) | 1.0 - 5.0 x 10^2 TCID50/mL | 5.0 - 10.0 x 10^2 TCID50/mL (generally less sensitive than N) |
| Cross-Reactivity Concern | Possible with other human coronaviruses (e.g., HKU1, OC43) | Lower, but variant-specific mAbs may lose binding |
| Best Use Case | High-sensitivity rapid tests for early infection | Tests differentiating infection from vaccination (if vaccine targets S only); entry inhibition studies |
Data sourced from current literature (2023-2024). TCID50: Median Tissue Culture Infectious Dose.
Detailed Experimental Protocols
Protocol: Standard Sandwich ELISA for Anti-SARS-CoV-2 Antibody Detection (Using Recombinant N/S Antigen)
Objective: To quantify IgG/IgM antibodies against SARS-CoV-2 N or S protein in human serum.
Materials:
- Coating Antigen: Recombinant SARS-CoV-2 N protein or S1/RBD protein.
- Microtiter Plate: 96-well, high-binding polystyrene.
- Blocking Buffer: 5% non-fat dry milk or 3% BSA in PBS-T (PBS + 0.05% Tween-20).
- Diluent: 1% BSA in PBS-T.
- Samples: Human serum/plasma (heat-inactivated at 56°C for 30 min).
- Detection Antibodies: HRP-conjugated goat anti-human IgG (γ-chain specific) and anti-human IgM (μ-chain specific).
- Substrate: TMB (3,3',5,5'-Tetramethylbenzidine).
- Stop Solution: 1M H2SO4.
- Plate Reader: For measuring absorbance at 450 nm.
Procedure:
- Coating: Dilute recombinant antigen to 1-2 µg/mL in carbonate-bicarbonate coating buffer (pH 9.6). Add 100 µL/well. Seal plate and incubate overnight at 4°C.
- Washing: Aspirate solution. Wash plate 3 times with 300 µL/well PBS-T using an automated or manual washer. Blot dry.
- Blocking: Add 300 µL/well blocking buffer. Incubate for 1-2 hours at room temperature (RT). Wash 3 times as in step 2.
- Sample Incubation: Prepare serial dilutions of serum samples in diluent (e.g., 1:50, 1:200, 1:800). Add 100 µL of diluted sample or diluent blank to designated wells. Incubate for 1-2 hours at RT. Wash 3 times.
- Detection Antibody Incubation: Dilute HRP-conjugated anti-human IgG or IgM in diluent per manufacturer's recommendation. Add 100 µL/well. Incubate for 1 hour at RT in the dark. Wash 5 times thoroughly.
- Signal Development: Add 100 µL/well TMB substrate. Incubate for 10-15 minutes at RT in the dark.
- Stop & Read: Add 50 µL/well 1M H2SO4 to stop reaction. Measure absorbance at 450 nm within 30 minutes.
- Analysis: Calculate cut-off value (typically mean of negative controls + 3 standard deviations). Samples with absorbance above the cut-off are considered positive.
Protocol: In-House Validation of a Rapid LFA for N Protein Detection
Objective: To determine the clinical sensitivity and specificity of a prototype SARS-CoV-2 N protein LFA.
Materials:
- Prototype LFA Cassettes.
- Clinical Samples: Nasopharyngeal swab samples in viral transport medium (VTM), previously characterized by RT-qPCR (positive and negative).
- VTM Buffer.
- Timer.
Procedure:
- Sample Preparation: Allow frozen VTM samples to thaw completely and vortex. For direct testing, use as is. Some protocols may involve a 1:1 dilution with the provided assay buffer.
- Test Execution: Remove cassette from pouch. Using the provided transfer pipette, add 3-4 drops (approx. 100 µL) of prepared sample to the sample well (S).
- Incubation: Start timer. Allow the test to develop at room temperature for exactly 15 minutes. Do not read results before or after this window.
- Result Interpretation: Visually inspect the cassette.
- Positive: Both control (C) and test (T) lines are visible, regardless of T line intensity.
- Negative: Only the control (C) line is visible.
- Invalid: Control (C) line does not appear. Repeat test with a new cassette.
- Data Collection: Record result for each sample. Compare to RT-qPCR gold standard.
- Analysis: Calculate Sensitivity = (True Positives / (True Positives + False Negatives)) x 100. Calculate Specificity = (True Negatives / (True Negatives + False Positives)) x 100.
Diagram 2: Immunoassay Sandwich Logic (87 chars)
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for SARS-CoV-2 Antigen Immunoassay Development
| Reagent / Material | Function & Purpose | Example/Typical Source |
|---|---|---|
| Recombinant N Protein | High-purity antigen for assay calibration, coating (ELISA), and antibody screening. | HEK293 or E. coli expressed, full-length or fragments. |
| Recombinant S1/RBD Protein | Antigen for S-targeted assays, neutralization assay development, and vaccine research. | Mammalian cell-expressed, glycosylated protein. |
| Monoclonal Anti-N Antibodies | Paired, non-competing antibodies for sandwich ELISA and LFA development. Essential for defining epitopes. | Mouse or rabbit hybridomas; must target distinct epitopes. |
| Monoclonal Anti-S/RBD Antibodies | Critical for variant analysis, neutralizing antibody detection, and S-protein capture assays. | Includes clones like CR3022 or variant-specific antibodies. |
| Colloidal Gold Nanoparticles (40nm) | The most common label for rapid LFAs. Antibodies are conjugated to the gold surface. | Commercially available as ready-to-conjugate suspensions. |
| HRP (Horseradish Peroxidase) | Enzyme label for high-sensitivity colorimetric (ELISA) or chemiluminescent assays. | Conjugated to secondary antibodies for detection. |
| Nitrocellulose Membrane | Porous matrix for capillary flow and immobilization of capture lines in LFAs. | Various pore sizes (e.g., 8-15 µm) to optimize flow and binding. |
| High-Binding ELISA Plates | Solid phase for immobilizing antigens or antibodies in plate-based assays. | Polystyrene plates with special surface treatment. |
| Pseudotyped VSV Particles | Safe, BSL-2 surrogate for live virus in neutralization assays targeting S protein entry. | Particles bearing SARS-CoV-2 S protein and a reporter gene (e.g., Luciferase). |
| Clinical Sample Panels | Well-characterized positive/negative human samples for clinical validation of assays. | Sourced from commercial biorepositories or collaborations. |
Within the broader thesis on SARS-CoV-2 genome structure and protein targets for diagnostics research, the need for rapid, sensitive, and specific nucleic acid detection platforms is paramount. CRISPR-Cas systems, originally characterized as adaptive immune mechanisms in bacteria, have been repurposed into powerful in vitro diagnostic tools. The SHERLOCK (Specific High-sensitivity Enzymatic Reporter unLOCKing) and DETECTR (DNA Endonuclease-Targeted CRISPR Trans Reporter) platforms represent groundbreaking approaches that leverage the programmable, sequence-specific recognition and collateral cleavage activities of certain Cas enzymes. This guide provides a technical deep dive into these platforms, with a focus on their application for SARS-CoV-2 detection.
Both platforms utilize Cas effector proteins that, upon recognition of their specific target nucleic acid sequence, exhibit collateral trans-cleavage activity, degrading nearby reporter molecules to generate a detectable signal.
SHERLOCK primarily employs Cas13a (from Leptotrichia wadei; LwCas13a) or Cas13b, which target single-stranded RNA (ssRNA). Cas13's collateral RNase activity cleaves fluorescent RNA reporters. DETECTR employs Cas12a (from Lachnospiraceae bacterium; LbCas12a), which targets double-stranded DNA (dsDNA). Cas12a's collateral DNase activity cleaves fluorescent DNA reporters.
The table below summarizes the core characteristics of each system.
Table 1: Core Comparison of SHERLOCK and DETECTR Platforms
| Feature | SHERLOCK (Cas13-based) | DETECTR (Cas12-based) |
|---|---|---|
| Primary Cas Enzyme | Cas13a (LwCas13a) or Cas13b | Cas12a (LbCas12a or AsCas12a) |
| Native Target | Single-stranded RNA (ssRNA) | Double-stranded DNA (dsDNA) |
| Collateral Activity | Nonspecific trans-cleavage of ssRNA | Nonspecific trans-cleavage of ssDNA |
| PAM/PFS Requirement | Protospacer Flanking Site (PFS) for Cas13a (e.g., non-G 3' of target); varies. | Protospacer Adjacent Motif (PAM) for Cas12a (e.g., TTTV 5' of target). |
| Typical Pre-amplification | RPA (Recombinase Polymerase Amplification) or RT-RPA | RPA or PCR |
| Reporter Molecule | Fluorescently quenched ssRNA probe (e.g., FAM-rU-rU-rU-BHQ-1) | Fluorescently quenched ssDNA probe (e.g., FAM-TTATT-BHQ-1) |
| Readout | Fluorescence (lateral flow strips or fluorometers) | Fluorescence (lateral flow strips or fluorometers) |
| Key SARS-CoV-2 Targets | N, E, S, Orf1ab genes | N, E genes |
Quantitative Performance Data
Sensitivity and specificity are critical for diagnostic applications. The following table consolidates key performance metrics from seminal and recent studies for SARS-CoV-2 detection.
Table 2: Reported Performance Metrics for SARS-CoV-2 Detection
| Platform / Assay Name | Limit of Detection (LoD) | Time-to-Result | Specificity | Clinical Sensitivity | Reference (Example) |
|---|---|---|---|---|---|
| SHERLOCK (v1, RT-RPA) | ~10-100 copies/µL | ~60 minutes | 100% (in silico) | 96% (vs. RT-PCR) | Science 2020 |
| SHERLOCK (with HUDSON) | ~6-125 copies/µL | <60 min | 100% | 90-100% | Nat. Biomed. Eng. 2020 |
| DETECTR (RT-LAMP) | 10 copies/µL | ~30-45 minutes | 100% (no cross-reactivity) | 95% (Positive Agreement) | Nat. Biotechnol. 2020 |
| STOPCovid (SHERLOCK variant) | ~100 copies/mL | ~60 min | 99.5% | 93.1% | N Engl J Med 2020 |
Detailed Experimental Protocols
Protocol 1: SHERLOCK-based SARS-CoV-2 RNA Detection
Principle: Viral RNA is converted to DNA via reverse transcription, amplified by RPA, then transcribed back to RNA by T7 RNA polymerase. The amplified RNA target activates LwCas13a, leading to collateral cleavage of an RNA reporter and fluorescence generation.
Materials (Research Reagent Solutions):
- LwCas13a Protein: Purified recombinant protein, the core collateral RNase effector.
- crRNA: Custom-designed, in vitro transcribed guide RNA targeting SARS-CoV-2 sequences (e.g., N gene).
- RT-RPA Master Mix: Contains reverse transcriptase, recombinase, polymerase, and nucleotides for isothermal amplification.
- T7 RNA Polymerase: For transcribing RPA amplicons into RNA.
- Fluorescent RNA Reporter: ssRNA oligo with 5'-FAM fluorophore and 3'-BHQ1 quencher (e.g., 5'-FAM-UU UU UU-BHQ1-3').
- Nucleoside Triphosphates (NTPs): For transcription and Cas13 reaction.
- Buffer System: Includes HEPES, MgCl2, DTT for optimal Cas13 activity.
Procedure:
- Sample Preparation & Lysis: Use heat, chemical lysis (e.g., HUDSON protocol), or column-based methods to extract RNA from nasopharyngeal swabs.
- Reverse Transcription & RPA Amplification:
- Combine 2 µL of extracted RNA with 0.5 µM forward/reverse RPA primers (containing T7 promoter on one primer), RT-RPA master mix, and nuclease-free water to 50 µL.
- Incubate at 37-42°C for 15-25 minutes.
- T7 Transcription & Cas13 Detection:
- Prepare a detection mix containing: 50-100 nM LwCas13a, 50-100 nM specific crRNA, 100 nM RNA reporter, 1 U/µL T7 RNA polymerase, 1 mM each NTP, and reaction buffer.
- Combine 5 µL of the RPA product with 15 µL of the detection mix.
- Incubate at 37°C for 10-60 minutes in a plate reader or heat block.
- Signal Readout:
- Measure real-time or endpoint fluorescence (Ex/Em: 485/535 nm for FAM).
- Lateral Flow Alternative: Use a biotinylated ssRNA reporter and FAM-labeled detection probe. Apply reaction to a strip with anti-FAM antibodies at the test line. Collateral cleavage prevents test line capture, causing signal inversion (cleavage = test line negative, control line positive).
Protocol 2: DETECTR-based SARS-CoV-2 DNA Detection
Principle: Viral RNA is reverse transcribed to DNA, amplified isothermally via RT-LAMP or RT-RPA. The dsDNA amplicon activates LbCas12a, triggering collateral cleavage of a DNA reporter.
Materials (Research Reagent Solutions):
- LbCas12a Protein: Purified recombinant protein, the core collateral DNase effector.
- crRNA: Designed to target the amplified SARS-CoV-2 DNA sequence, considering the TTTV PAM.
- RT-LAMP/RPA Master Mix: Contains Bst polymerase, reverse transcriptase, and primers for isothermal amplification.
- Fluorescent ssDNA Reporter: e.g., 5'-6-FAM-TTATT-3IABkFQ-3' (IABkFQ is a quencher).
- Nucleoside Triphosphates (dNTPs): For amplification.
- Reaction Buffer: Typically includes Tris-HCl, MgCl2, DTT.
Procedure:
- Sample Preparation: Viral RNA extraction as above.
- Reverse Transcription & Isothermal Amplification (RT-LAMP):
- Combine RNA with RT-LAMP master mix containing 6-8 primers targeting SARS-CoV-2, dNTPs, and Bst polymerase.
- Incubate at 60-65°C for 20-30 minutes.
- Cas12 Detection:
- Prepare a detection mix containing: 50-100 nM LbCas12a, 50-100 nM specific crRNA, 100-500 nM ssDNA reporter in reaction buffer.
- Combine a small aliquot (2-5 µL) of the LAMP product with the detection mix.
- Incubate at 37°C for 5-15 minutes.
- Signal Readout:
- Measure fluorescence (Ex/Em: 485/535 nm). Signal generation is rapid upon target recognition.
- Lateral Flow: Use a FAM-labeled ssDNA reporter and biotinylated ssDNA oligo. Uncleaved reporter forms a complex with anti-FAM gold nanoparticles and is captured by biotin at the test line. Cleavage eliminates the test line signal.
Visualizations of Workflows and Mechanisms
Diagram 1: SHERLOCK Assay Workflow for SARS-CoV-2 RNA
Diagram 2: DETECTR Assay Workflow for SARS-CoV-2
Diagram 3: CRISPR-Cas Collateral Cleavage Signal Generation
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for CRISPR-Based Diagnostics
| Reagent | Function in Assay | Example (SARS-CoV-2 Context) | Critical Parameters |
|---|---|---|---|
| Recombinant Cas Protein | Core effector enzyme with collateral nuclease activity. | LwCas13a (SHERLOCK) or LbCas12a (DETECTR). | Purity, concentration, storage buffer (glycerol, salt), nuclease contamination. |
| Synthetic crRNA | Provides sequence specificity by guiding Cas to target. | 20-30 nt spacer complementary to N gene, plus Cas-specific direct repeat. | Design (avoid off-targets, secondary structure), chemical modification (stability), synthesis scale. |
| Isothermal Amplification Mix | Amplifies target from low copy numbers without thermocycler. | RT-RPA or RT-LAMP kits with reverse transcriptase. | Speed, efficiency, tolerance to inhibitors, primer design complexity. |
| Fluorescent Quenched Reporter | Signal-generating substrate cleaved during collateral activity. | FAM-UUUUUU-BHQ1 (RNA) or FAM-TTATT-IABkFQ (DNA). | Quencher efficiency (BHQ1, IABkFQ), linkage chemistry, solubility. |
| NTPs/dNTPs | Substrates for amplification and transcription steps. | Pure mixes of ATP, CTP, GTP, UTP or dNTPs. | Concentration, purity (free from RNase/DNase), pH. |
| Optimized Reaction Buffer | Maintains optimal pH, salt, and cofactor conditions. | Contains HEPES, MgCl2, DTT, PEG, etc. | Mg2+ concentration (critical for Cas activity), reducing agents, crowding agents. |
| Lateral Flow Strips | For visual, instrument-free readout. | Strip with anti-FAM test line and control line. | Membrane type (nitrocellulose), conjugate pad composition, capillary flow time. |
This technical guide is framed within a broader thesis on the SARS-CoV-2 genome structure and protein targets for diagnostics research. The SARS-CoV-2 genome, approximately 30 kb in length, encodes four structural proteins (Spike (S), Envelope (E), Membrane (M), and Nucleocapsid (N)), sixteen non-structural proteins (nsp1-16), and several accessory proteins. This genomic complexity offers multiple targets for diagnostic assay development. The central thesis posits that leveraging this multi-target architecture through multiplex assays is critical for overcoming limitations of single-target tests—such as false negatives due to viral mutation or low viral load—thereby achieving unprecedented diagnostic accuracy, robustness, and confidence in pandemic and post-pandemic surveillance.
Core Principles and Advantages of Multiplexing
Multiplex assays concurrently detect two or more distinct viral targets within a single reaction. For SARS-CoV-2, this typically involves targeting conserved regions across multiple genes.
Key Advantages:
- Enhanced Accuracy & Reduced False Negatives: Targeting multiple genes mitigates the impact of point mutations, deletions, or variations in gene expression that could impair detection of a single target.
- Strain Differentiation and VOC Tracking: Designed with targets in variable regions, multiplex assays can distinguish between variants of concern (VOCs) in a single test.
- Comprehensive Pathogen Detection: Can be expanded to co-detection of SARS-CoV-2 with other respiratory viruses (e.g., influenza, RSV), aiding differential diagnosis.
- Efficiency: Conserves precious sample material and reduces reagent costs and hands-on time per data point.
Quantitative Data on SARS-CoV-2 Genomic Targets
Table 1: Key SARS-CoV-2 Genomic Targets for Multiplex Assay Design
| Target Gene/Region | Genomic Position (approx.) | Function | Relative Abundance in Viral Transcriptome | Conservation | Suitability for Diagnostics |
|---|---|---|---|---|---|
| N gene (Nucleocapsid) | 28274-29533 | RNA packaging, virion assembly | High; abundantly expressed subgenomic RNA | High | Excellent primary target; highly expressed and conserved. |
| E gene (Envelope) | 26245-26472 | Virion assembly and budding | Moderate | Very High | Excellent confirmatory target; small and highly conserved. |
| RdRp gene (in ORF1ab) | 13442-13468 | Viral RNA replication | Low (genomic RNA only) | Extremely High | Excellent confirmatory target; part of the replicase complex, highly conserved but less abundant. |
| S gene (Spike) | 21563-25384 | Host cell receptor binding, fusion | Moderate | Moderate-Low (subject to mutation) | Useful for variant discrimination; mutations common (e.g., Delta, Omicron deletions). |
| ORF1ab (non-RdRp) | Various | Viral replication/transcription complex | Low (genomic RNA only) | High | Good target, but sequence complexity requires careful design. |
Table 2: Performance Comparison of Representative Multiplex Assays
| Assay Platform | Targets Detected (SARS-CoV-2) | Additional Pathogens | Reported Clinical Sensitivity | Reported Clinical Specificity | Turnaround Time |
|---|---|---|---|---|---|
| CDC 2019-nCoV RT-PCR Panel | N1, N2 | None | >99% (for N1/N2) | 100% | ~4 hours |
| Multiplex RT-qPCR (Research Assay) | N gene, E gene, RdRp | None | 98.5% (vs. single-target 95%) | 100% | ~2 hours |
| Commercial Multiplex PCR Panel | N gene, S gene target | Influenza A/B, RSV | 99.2% for SARS-CoV-2 | 99.9% for all targets | ~1.5 hours |
| CRISPR-based Multiplex (e.g., SHERLOCK) | N gene, Orf1ab | None | 97% | 100% | ~1 hour |
Detailed Experimental Protocols
Protocol: Design and Validation of a Triplex RT-qPCR Assay for SARS-CoV-2
Objective: To simultaneously detect three conserved SARS-CoV-2 targets (N, E, RdRp) in a single-tube, single-channel (SYBR Green) or multi-channel (TaqMan probe) RT-qPCR reaction.
Materials: See "The Scientist's Toolkit" below.
Methodology:
A. Primer/Probe Design:
- Sequence Alignment: Retrieve complete SARS-CoV-2 genomes from GISAID or NCBI. Perform multiple sequence alignment using software (e.g., Clustal Omega, Geneious) focusing on conserved regions across N, E, and ORF1ab (RdRp) genes.
- In Silico Validation: Use tools like Primer-BLAST to ensure specificity against the human genome and other coronaviruses. Check for secondary structure and dimer formation.
- Dye/Quencher Selection (for TaqMan): Label probes with distinct fluorophores (e.g., FAM for N gene, HEX/VIC for E gene, Cy5 for RdRp) matched to your qPCR instrument's channels. Use compatible quenchers (e.g., BHQ-1, BHQ-2).
B. Assay Optimization:
- Master Mix Preparation: Use a robust one-step RT-qPCR master mix suitable for multiplexing.
- Primer/Probe Concentration Gradient: Perform a matrix titration (e.g., 50-900 nM for primers, 50-250 nM for probes) to find concentrations that yield the lowest Cq (quantification cycle) values and highest fluorescence amplitude without cross-talk or inhibition.
- Thermal Cycling Conditions: Standard conditions: Reverse Transcription: 50°C for 10-15 min; Initial Denaturation: 95°C for 2 min; 45 cycles of: Denaturation 95°C for 5-10 sec, Annealing/Extension 60°C for 30-45 sec (acquire fluorescence). Annealing temperature may require optimization (± 3°C).
C. Validation:
- Analytical Sensitivity (Limit of Detection - LoD): Test serial dilutions of SARS-CoV-2 RNA (characterized standard). The LoD is the lowest concentration detected in ≥95% of replicates. Perform for each target individually and in multiplex.
- Analytical Specificity: Test against RNA/DNA from other human coronaviruses (HKU1, OC43, etc.), influenza, RSV, and human genomic DNA to confirm no cross-reactivity.
- Clinical Validation: Run assay on a panel of confirmed positive (various Ct values) and negative clinical nasopharyngeal swab extracts. Compare results to a gold-standard singleplex assay.
Protocol: High-Throughput Multiplex NGS for Strain Surveillance
Objective: To amplify and sequence multiple regions of the SARS-CoV-2 genome for variant identification and detection of co-infections.
Methodology:
- Multiplex Primer Panel (Amplicon Sequencing): Use a tiled, multiplex primer scheme (e.g., ARTIC Network protocol) to generate ~400 bp overlapping amplicons covering the entire genome.
- Library Preparation: Perform one-step RT-PCR with multiplex primer pools. Clean up amplicons. Attach dual indices and sequencing adapters via a secondary limited-cycle PCR.
- Bioinformatic Analysis: Demultiplex reads by sample and primer pair. Map reads to a reference genome (MN908947.3). Call variants and assign lineage using tools like Pangolin. Analyze for mixed populations indicating potential co-infection.
Visualizations
Workflow for Multiplex RT-qPCR Detection
SARS-CoV-2 Genome with Key Diagnostic Targets
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for Multiplex Assay Development
| Reagent/Material | Function & Importance in Multiplexing | Example/Note |
|---|---|---|
| One-Step RT-qPCR Master Mix | Integrates reverse transcription and PCR amplification in a single buffered solution. Critical for workflow simplicity and reducing contamination. Must be optimized for multiplexing (high processivity, inhibitor tolerance). | TaqMan Fast Virus 1-Step, qScript XLT One-Step. |
| Sequence-Specific Oligonucleotides | Primers and probes for each target. Probes require distinct fluorophore/quencher pairs. HPLC or equivalent purification is essential for performance. | Primers (~20-25 bp), Dual-labeled hydrolysis probes (e.g., FAM/BHQ-1, HEX/BHQ-1, Cy5/BHQ-2). |
| SARS-CoV-2 RNA Standards | Quantitated, full-genome or target-specific RNA for assay optimization, determining LoD, and creating standard curves. | Available from BEI Resources, ATCC, or commercial IVD manufacturers. |
| Nuclease-Free Water & Plasticware | To prevent degradation of RNA and oligonucleotides. Pipette tips with filters are recommended. | Certified RNase/DNase-free. |
| Multi-Channel Real-Time PCR Instrument | Instrument capable of exciting and detecting multiple fluorescent dyes simultaneously across 96- or 384-well plates. | Applied Biosystems QuantStudio, Bio-Rad CFX, Roche LightCycler 480 II. |
| Negative & Inhibition Controls | Human genomic DNA, other respiratory pathogen RNA, and internal control (e.g., RNase P) to monitor sample quality and PCR inhibition. | Critical for validating specificity and assay robustness. |
| Bioinformatics Software | For primer design, sequence alignment, and analysis of NGS-based multiplex assays. | Geneious, Primer-BLAST, Galaxy Platform, Artic Network pipelines. |
Optimizing Diagnostic Assays: Troubleshooting Primer Mismatches, Antic Escape, and Sensitivity Issues
This whitepaper is framed within a broader thesis investigating the SARS-CoV-2 genome structure and protein targets for diagnostic research. A primary challenge for molecular diagnostics is the virus's rapid evolution, leading to mutations that degrade primer/probe binding efficacy. This guide details in silico strategies to design resilient assays and establish rapid update protocols to maintain diagnostic accuracy.
The Mutation Challenge: Quantitative Impact on Assay Performance
Table 1: Impact of Key SARS-CoV-2 Variant Mutations on Common Diagnostic Assay Targets (2023-2024 Data)
| WHO Variant Designation | Key Spike Mutations | Common qPCR Target Genes Affected | Reported Impact on Assay Efficiency (Ct Shift) | Prevalence (%) in GISAID (as of Q1 2024) |
|---|---|---|---|---|
| Omicron JN.1 | L455S, F456L | S-gene (several assays) | ΔCt +2.5 to >7.0 (S-dropout common) | ~85% |
| Omicron XBB.1.5 | F486P, R493Q | S-gene, ORF1a | ΔCt +1.5 to +3.0 | <5% |
| BA.5.2 | del69-70, R346T | S-gene (TaqPath dropout) | ΔCt >5.0 (characteristic dropout) | <1% |
| Delta (B.1.617.2) | L452R, T478K | S-gene | ΔCt +1.0 to +2.5 | Negligible |
Data synthesized from recent publications in Eurosurveillance, GISAID metadata, and FDA Molecular Diagnostic Templates.
In SilicoPrimer/Probe Design Workflow
Core Algorithmic Strategy
The design pipeline prioritizes conserved genomic regions identified through entropy analysis of aligned SARS-CoV-2 sequences.
Experimental Protocol 1: Conserved Region Identification
- Input: Download ≥10,000 representative complete SARS-CoV-2 genomes from GISAID (prioritizing last 6 months).
- Alignment: Use MAFFT v7.520 for multiple sequence alignment against reference NC_045512.2.
- Entropy Calculation: Compute Shannon entropy (H) for each nucleotide position (i) across alignment (N sequences):
H(i) = -Σ (p_x * log2(p_x)) for x ∈ {A, T, C, G}, wherep_xis frequency of base x. - Output: Genomic regions with H < 0.2 over a window of ≥50 bases are flagged as high-conservation targets.
Diagram 1: In Silico Assay Design and Validation Workflow
Design Parameters for Mutation Resilience
- Length: Primers 18-25 bp; Probes 15-20 bp.
- Tm: Primer Tm 55-60°C; Probe Tm 7-10°C higher.
- 3'-End Stability: Avoid G/C clamps at 3'-end to reduce mis-priming on mismatched templates.
- Degeneracy & Wobble Bases: Strategically incorporate inosine or universal bases (e.g., locked nucleic acids - LNAs) at known variable positions while maintaining overall Tm.
Experimental Protocol forIn VitroValidation
Experimental Protocol 2: In Vitro Cross-reactivity and Limit of Detection (LOD) Testing
- Synthetic Controls: Obtain synthetic RNA controls (Twist Bioscience) for wild-type and variant sequences (including mismatches).
- qPCR Setup: Use AgPath-ID One-Step RT-PCR Kit. Reaction: 5 µL RNA, 10 µL 2x buffer, 1 µL enzyme, 0.5 µL each primer/probe (final conc. 400 nM/200 nM), nuclease-free water to 20 µL.
- Thermocycling: 50°C for 15 min (RT); 95°C for 2 min; 45 cycles of 95°C for 3s, 60°C for 30s (acquire fluorescence).
- Analysis: Calculate LOD via probit analysis on 8 replicates of serial 10-fold dilutions (10^6 to 10^0 copies/µL). Test cross-reactivity against human genomic DNA and common respiratory pathogen panels.
Table 2: Key Research Reagent Solutions for Validation
| Reagent / Material | Manufacturer / Source | Function in Protocol |
|---|---|---|
| Synthetic SARS-CoV-2 RNA Controls | Twist Bioscience, ATCC | Provides stable, quantifiable templates for wild-type and variant sequences, essential for LOD and mismatch testing. |
| AgPath-ID One-Step RT-PCR Kit | Thermo Fisher Scientific | Integrated reverse transcription and qPCR chemistry for streamlined one-tube assay validation. |
| QuantStudio 7 Pro Real-Time PCR System | Thermo Fisher Scientific | High-throughput thermocycler with multiplexing capability for testing multiple primer sets simultaneously. |
| Human Genomic DNA | Promega, Roche | Negative control template to assess primer/probe specificity and rule off-target amplification. |
| Respiratory Pathogen Panel RNA | ZeptoMetrix, Bio-Rad | Contains RNA/DNA from common coronaviruses, influenza, RSV, etc., to validate diagnostic specificity. |
| Nuclease-Free Water (Molecular Grade) | Ambion, Sigma-Aldrich | Critical for preventing RNase/DNase degradation of sensitive oligonucleotides and RNA templates. |
Update Strategy: A Continuous Monitoring Framework
Diagram 2: Continuous Assay Monitoring & Update Trigger Logic
Update Protocol: When the logic in Diagram 2 triggers a review, the design pipeline (Diagram 1) is initiated. The updated candidate assay must pass in silico specificity and in vitro LOD validation against both the emergent variant and prior circulating strains before deployment, ensuring backward compatibility.
Integrating systematic in silico design focused on low-entropy genomic regions with a robust, data-triggered update strategy is critical for maintaining diagnostic accuracy against evolving SARS-CoV-2. This approach, rooted in a deep understanding of viral genome structure, provides a model for resilient molecular diagnostic development against any RNA virus.
1. Introduction: Evolving Targets in a Broader Genomic Context The effectiveness of rapid antigen tests (RATs) for SARS-CoV-2 is contingent on the high-affinity binding of monoclonal antibodies (mAbs) to conserved viral nucleocapsid (N) protein epitopes. This discussion is framed within a broader thesis on SARS-CoV-2 genome structure, which reveals that diagnostic and therapeutic targets are under differential evolutionary pressure. While spike (S) protein mutations drive immune evasion, mutations in the N protein—a critical diagnostic target—can arise from host immune pressure or genetic hitchhiking, compromising test sensitivity. The emergence of the Omicron lineage (BA.1, BA.2, BA.5, and subsequent variants like XBB and JN.1) marked a pivotal shift, with numerous mutations in the N gene leading to altered protein expression and conformation, posing significant challenges for N-protein-dependent diagnostics.
2. Quantitative Analysis of N Protein Mutations and Test Performance Impact The following tables summarize key mutations and their demonstrated impact on antigen test performance.
Table 1: Prevalent N Protein Mutations in Omicron and Subsequent Variants
| Variant Lineage | Key N Protein Mutations (Relative to Wuhan-Hu-1) | Structural/Functional Domain |
|---|---|---|
| Omicron BA.1 | P13L, Δ31-33, R203K, G204R | NTD, Linker Region |
| Omicron BA.2 | P13L, Δ31-33, R203K, G204R, S413R | NTD, Linker, CTD |
| Omicron BA.5 | P13L, Δ31-33, R203K, G204R, S413R | NTD, Linker, CTD |
| XBB.1.5 | P13L, Δ31-33, R203K, G204R, S413R | NTD, Linker, CTD |
| JN.1 | P13L, Δ31-33, R203K, G204R, S413R, L335S | NTD, Linker, CTD |
Table 2: Documented Impact on Commercial Antigen Test Sensitivity
| Study (Example) | Variant Tested | Test(s) Evaluated | Reported Sensitivity Reduction (vs. Ancestral) | Putative Cause |
|---|---|---|---|---|
| Bekliz et al., 2022 | BA.1 | 3 commercial kits | Up to 33% reduction | R203K/G204R altering mAb affinity |
| Rönnberg et al., 2022 | BA.1, BA.2 | 5 commercial kits | Variable; some tests unaffected | Epitope disruption by Δ31-33 & P13L |
| FDA Monograph Data | BA.5, XBB | Multiple EUA tests | Generally maintained; kit-specific failures noted | Conformational changes from cumulative mutations |
3. Experimental Protocols for Evaluating Antigen Test Efficacy
Protocol 1: In Vitro Assessment of mAb Binding Affinity Using ELISA Objective: Quantify the binding affinity of diagnostic mAbs to recombinant wild-type and variant N proteins. Materials: 96-well plates coated with recombinant N protein variants, blocking buffer (5% BSA/PBS), primary diagnostic mAb (from test strip), HRP-conjugated secondary antibody, TMB substrate, stop solution (1M H2SO4). Procedure:
- Coat plates with 100 µL/well of 2 µg/mL recombinant N protein (ancestral and variant) in PBS overnight at 4°C.
- Wash 3x with PBS-T (0.05% Tween-20). Block with 200 µL/well blocking buffer for 1 hour at room temperature (RT).
- Wash 3x. Add 100 µL/well of serially diluted primary mAb in PBS. Incubate 2 hours at RT.
- Wash 3x. Add 100 µL/well of appropriate HRP-conjugated secondary antibody (1:5000). Incubate 1 hour at RT.
- Wash 5x. Develop with 100 µL/well TMB substrate for 15 minutes in the dark.
- Stop reaction with 50 µL/well stop solution. Measure absorbance at 450 nm immediately.
- Calculate EC50 values using a 4-parameter logistic curve to compare binding affinity.
Protocol 2: Testing Clinical Sensitivity with Live Virus Isolates Objective: Determine the limit of detection (LoD) of an antigen test for different live variants. Materials: Vero E6/TMPRSS2 cells, authentic virus isolates (ancestral, Delta, Omicron sub-variants), viral transport media, serial dilution series, commercial antigen test kits, qRT-PCR for TCID50 correlation. Procedure:
- Propagate and titrate virus stocks to determine TCID50/mL.
- Prepare 10-fold serial dilutions of each virus in viral transport media, covering a range from 10^5 to 10^1 TCID50/mL.
- For each dilution, apply 100 µL to the sample pad of the antigen test according to the manufacturer's instructions. Perform tests in triplicate.
- Develop for the exact time specified. Record visual results and use a lateral flow reader for quantitative band intensity.
- Determine the LoD as the lowest viral concentration yielding a positive result in ≥95% of replicates.
- Parallel test all samples with qRT-PCR to correlate LoD with RNA copy number.
4. Visualization of Diagnostic Target Disruption and Validation Workflow
Diagram Title: N Protein Mutation Impact on Diagnostic Efficacy
Diagram Title: Antigen Test Validation Workflow for Variants
5. The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Antigen Test Optimization Research
| Reagent / Material | Function / Rationale | Example Source / Note |
|---|---|---|
| Recombinant N Protein (Variants) | Key substrate for initial binding studies; must include Omicron-specific mutations (P13L, Δ31-33, R203K/G204R). | Commercial suppliers (e.g., Sino Biological, Acro Biosystems); ensure full-length and proper folding. |
| Diagnostic mAb Clones (Paired) | Capture and detection antibodies from commercial test kits; essential for epitope mapping and affinity measurements. | Isolated from hybridomas or obtained via commercial partnership/custom synthesis. |
| Authentic Virus Isolates (Variants) | Gold standard for LoD studies; reflects native protein conformation and oligomerization. | Obtain from repositories (BEI Resources, ATCC) under appropriate biosafety (BSL-3) protocols. |
| Pseudotyped Virus Particles (N-Variant) | Safer (BSL-2) alternative for neutralization and some binding studies; incorporates variant N protein. | Custom production via lentiviral/vesicular stomatitis virus (VSV) backbone systems. |
| Vero E6 / TMPRSS2 Cells | Cell line for propagating authentic virus isolates and conducting virus neutralization tests. | Standard cell culture repositories; ensures high viral titers. |
| Lateral Flow Test Strips (Bare) | Unconjugated nitrocellulose membranes and pads for prototyping new mAb pairs and optimizing test geometry. | Suppliers like MilliporeSigma, GE Healthcare. |
| Surface Plasmon Resonance (SPR) Chip | For real-time, label-free kinetic analysis (KD, Kon, Koff) of mAb-antigen interactions with high precision. | Instruments like Biacore; CMS sensor chips. |
Mitigating Cross-Reactivity with Endogenous Human Coronaviruses (HCoVs) and Other Pathogens
Within the imperative to develop precise SARS-CoV-2 diagnostics, cross-reactivity with antigenically related endemic coronaviruses (HCoV-229E, -NL63, -OC43, -HKU1) and other common pathogens presents a significant challenge. This whitepaper provides a technical framework for the design and validation of molecular and immunoassays targeting unique genomic and structural features of SARS-CoV-2, thereby ensuring diagnostic specificity. The content is framed within a broader thesis on SARS-CoV-2 genome architecture and its exploitation for specific diagnostic target identification.
The SARS-CoV-2 genome shares significant homology with other members of the Coronaviridae family, particularly in conserved regions such as the RNA-dependent RNA polymerase (RdRp). Furthermore, structural proteins like the nucleocapsid (N) protein exhibit conserved epitopes. Antibodies elicited by prior HCoV infections or assays using conserved primers/probes can thus yield false-positive results. Mitigating this requires a detailed understanding of divergent genomic sequences and tertiary protein structures unique to SARS-CoV-2.
Comparative Genomic Analysis for Target Selection
The key to specific diagnostics lies in targeting genomic regions of high divergence. The following table summarizes quantitative homology data for key SARS-CoV-2 proteins versus endemic HCoVs, based on recent multiple sequence alignments (e.g., from NCBI Virus and GISAID).
Table 1: Protein Sequence Homology of SARS-CoV-2 vs. Endemic HCoVs
| SARS-CoV-2 Protein | % Identity to HCoV-OC43 | % Identity to HCoV-229E | % Identity to HCoV-NL63 | % Identity to HCoV-HKU1 | Recommended for Specific Targeting? |
|---|---|---|---|---|---|
| Spike (S) Glycoprotein | ~30% | ~28% | ~32% | ~31% | Yes (focus on S1/RBD) |
| Nucleocapsid (N) | ~48% | ~29% | ~36% | ~50% | Caution (requires divergent region) |
| Membrane (M) | ~39% | ~35% | ~37% | ~40% | Moderate |
| Envelope (E) | ~35% | ~25% | ~30% | ~37% | Yes |
| RNA-dependent RNA Polymerase (RdRp) | ~96% (nsp12) | ~67% | ~68% | ~95% | No (highly conserved) |
Note: Homology percentages are approximations from latest alignments; the RdRp is conserved for function, making it a poor discriminatory target.
Experimental Protocols for Validation
1In SilicoSpecificity Screening Protocol
Objective: Bioinformatic validation of primer/probe or epitope specificity. Methodology:
- Sequence Retrieval: Curate comprehensive databases of full-length genomes for SARS-CoV-2 variants and all endemic HCoVs, plus common respiratory pathogens (Influenza A/B, RSV, M. pneumoniae), from NCBI GenBank and GISAID.
- Target Alignment: Perform multiple sequence alignment (MSA) using MAFFT or Clustal Omega for the candidate target region (e.g., a 150-nucleotide window within the N gene).
- Specificity Check: Use BLASTN (for primers) or BLASTP (for epitopes) against the "nt" and "nr" databases, restricting to human pathogens.
- Mismatch Analysis: Manually inspect alignments for 3'-end mismatches in primer sequences (critical for PCR) and conserved linear/ conformational B-cell epitopes.
2In VitroCross-Reactivity Testing Protocol for Immunoassays
Objective: Empirically test monoclonal/polyclonal antibodies against recombinant proteins from related pathogens. Methodology:
- Antigen Coating: Coat ELISA plates with 100 ng/well of recombinant proteins: SARS-CoV-2 N, SARS-CoV-2 S1, and homologs from HCoV-OC43, HKU1, 229E, NL63. Include common viral antigens (e.g., Influenza NP).
- Antibody Incubation: Apply the candidate detection antibody (monoclonal) or serum (polyclonal) in serial dilutions (starting at 1 µg/mL for mAbs) in duplicate. Include a no-antigen control.
- Detection & Thresholding: Use standard colorimetric detection. Calculate cross-reactivity as: (Signal from heterologous antigen / Signal from homologous SARS-CoV-2 antigen) x 100%. A value >5% indicates significant cross-reactivity.
- Competitive Inhibition: Perform a competitive ELISA with pre-incubation of antibody with soluble heterologous antigens to confirm specificity.
Visualization of Diagnostic Development Workflow
Diagram 1: Diagnostic Specificity Development Workflow (98 chars)
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Cross-Reactivity Studies
| Reagent / Material | Supplier Examples | Function in Cross-Reactivity Mitigation |
|---|---|---|
| Recombinant SARS-CoV-2 Proteins (S1, RBD, N, M) | Sino Biological, RayBiotech, The Native Antigen Company | Positive control antigens for assay development and competitive inhibition studies. |
| Recombinant Endemic HCoV Proteins (OC43, HKU1, 229E, NL63) | Sino Biological, RayBiotech | Critical negative controls to empirically test antibody or assay cross-reactivity. |
| Synthetic SARS-CoV-2 RNA Genomes (Full-length or fragments) | Twist Bioscience, ATCC | Molecular standards for qPCR specificity testing, free of background pathogen co-infection. |
| Characterized Human Positive Serum Panels (Pre-pandemic & COVID-19 convalescent) | BEI Resources, SeraCare | Validate serological assay specificity against a background of endemic HCoV antibodies. |
| Pathogen Genomic DNA/RNA Panels (Respiratory Panel) | ATCC, ZeptoMetrix | Control templates for molecular diagnostic specificity testing against common co-circulating pathogens. |
| Cross-reactive Monoclonal Antibodies (e.g., anti-HCoV OC43 N protein) | Numerous academic sources; available via BEI | Tools to map conserved epitopes and validate blocking strategies in immunoassays. |
Advanced Strategies: Targeting Unique Structural Features
Beyond linear sequence, exploiting unique conformational epitopes in the Spike protein's receptor-binding motif (RBM) or quaternary structures of the nucleocapsid phosphoprotein oligomers offers high specificity. Cryo-EM and X-ray crystallography data of SARS-CoV-2 proteins are indispensable for this structure-guided design.
Diagram 2: Strategies to Overcome Cross-Reactivity (86 chars)
The development of specific SARS-CoV-2 diagnostics necessitates a multi-layered strategy that begins with intelligent target selection informed by comparative genomics and structural biology. Rigorous in silico and in vitro validation against a panel of endemic HCoVs and common pathogens is non-negotiable. By adhering to the protocols and utilizing the toolkit outlined herein, researchers can effectively mitigate cross-reactivity, ensuring diagnostic accuracy essential for public health and clinical decision-making.
Within the broader thesis on SARS-CoV-2 genome structure and its protein targets for diagnostics, optimizing sample collection is paramount. The viral load dynamics, influenced by the genomic regions targeted (e.g., ORF1ab, N, E, S genes), vary significantly across anatomical sites. This technical guide synthesizes current data and methodologies for three primary sample types, providing researchers with a framework to maximize nucleic acid yield and diagnostic accuracy for downstream genomic and proteomic analysis.
Comparative Viral Load Dynamics by Sample Type
Recent meta-analyses and cohort studies indicate consistent, though variable, relationships between sample type and detectable SARS-CoV-2 RNA. The following table summarizes key quantitative findings.
Table 1: Comparative Viral Load and Detection Sensitivity by Sample Type
| Sample Type | Median Viral Load (Log10 copies/mL) | Relative Sensitivity vs. NP (95% CI) | Optimal Collection Timing Post-Symptom Onset | Key Genomic Targets with Highest Yield |
|---|---|---|---|---|
| Nasopharyngeal (NP) Swab | 5.2 - 7.1 | 1.00 (Reference) | Days 1-5 | ORF1ab, N gene |
| Saliva (Deep-Cough) | 4.8 - 6.9 | 0.95 (0.90-0.98) | Days 1-7 | S gene, N gene |
| Anterior Nasal (AN) Swab | 4.5 - 6.3 | 0.88 (0.82-0.93) | Days 1-5 | N gene |
Data synthesized from live search results of recent studies (2023-2024). NP swabs remain the gold standard for peak viral load, but saliva shows comparable sensitivity with broader detection window and less variability in self-collection.
Detailed Experimental Protocols for Sample Processing
Protocol 1: Optimized RNA Extraction from Nasopharyngeal Swabs
This protocol is designed for maximum recovery of intact viral RNA for genome sequencing.
Materials: Viral Transport Medium (VTM), NP swabs (flocked nylon), magnetic bead-based RNA extraction kit, proteinase K, ethanol (100%, 70%), nuclease-free water.
Procedure:
- Elution: Vortex NP swab in 3 mL VTM for 30 seconds. Incubate at room temperature for 10 minutes.
- Lysis: Transfer 300 µL of VTM to a sterile tube. Add 300 µL of lysis buffer containing 5 µL proteinase K. Mix thoroughly and incubate at 56°C for 15 minutes.
- Binding: Add 600 µL of 100% ethanol to the lysate. Transfer the mixture to a magnetic bead column, incubate for 5 minutes, and apply a magnetic field. Discard flow-through.
- Washes: Perform two washes with 700 µL of wash buffer (containing ethanol). Perform a final wash with 700 µL of 70% ethanol.
- Elution: Dry the column for 5 minutes. Elute RNA in 60 µL of pre-heated (65°C) nuclease-free water. Store at -80°C.
Protocol 2: Saliva Sample Collection and Stabilization for Viral Load Quantification
This protocol addresses the inherent RNase activity in saliva, crucial for preserving viral RNA integrity.
Materials: Saliva collection aid (e.g., Oragene•RNA kit), RNA stabilization buffer, dry ice, centrifugation equipment.
Procedure:
- Collection: Instruct the donor to cough deeply 3-5 times prior to collection. Deposit 1-2 mL of saliva directly into a tube containing 1 mL of stabilization buffer. Cap and invert immediately.
- Homogenization: Vortex the sample for 20 seconds. Incubate the tube at 50°C for 1 hour to inactivate viral particles and nucleases.
- Clarification: Centrifuge at 4,500 x g for 20 minutes at 4°C. Carefully transfer the clarified supernatant to a fresh tube.
- Storage/Processing: Supernatant can be stored at -80°C or processed immediately using a high-volume (≥500 µL input) RNA extraction protocol, as described in Protocol 1.
Protocol 3: Anterior Nasal Swab Self-Collection and Processing
Optimized for at-home collection kits, focusing on user compliance and stability.
Materials: Sterile AN swabs (flocked or spun polyester), dry tube or minimal volume of stabilizing solution, buffer EL.
Procedure:
- Collection: Insert swab ~1-2 cm into one nostril, rotating against the nasal wall for 5 complete circles. Repeat in the other nostril with the same swab.
- Storage/Transport: Place swab immediately into a dry sterile tube or a tube containing 1 mL of a DNA/RNA shield buffer. The sample is stable at room temperature for up to 72 hours.
- Elution: Add 1 mL of buffer EL to the dry tube or use the provided liquid. Vortex vigorously for 1 minute. Incubate at room temperature for 10 minutes.
- Processing: Use 300 µL of the eluate as input for standard RNA extraction (see Protocol 1, Step 2 onwards).
Sample Type Decision Pathway for Diagnostic Research
Title: Sample Type Selection Logic for SARS-CoV-2 Research
SARS-CoV-2 Genomic Targets in Diagnostic Assay Development
The relationship between sample type, viral load, and the detectability of key genomic regions is critical for assay design.
Title: Sample-Type Efficacy for Key SARS-CoV-2 Genomic Targets
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 2: Key Reagents for Sample Collection, Processing, and Analysis
| Reagent/Material | Primary Function | Key Consideration for Optimization |
|---|---|---|
| Flocked Nylon Swabs (for NP/AN) | Maximizes cellular and viral particle absorption and elution. | Superior to spun polyester for RNA yield from NP samples. |
| Viral Transport Medium (VTM) | Preserves viral integrity during transport. | Must be validated for compatibility with downstream extraction kits. |
| DNA/RNA Shield or Similar Stabilization Buffer | Inactivates nucleases and stabilizes nucleic acids in saliva/AN swabs. | Enables room-temperature storage, critical for home-collection studies. |
| Magnetic Bead-Based RNA Extraction Kits | High-throughput, automated purification of viral RNA. | Select kits with larger input volumes (≥500 µL) for saliva supernatants. |
| Proteinase K | Digests proteins and inactivates nucleases during lysis. | Critical step for efficient release of viral RNA from capsid. |
| PCR Primers/Probes for Multi-Gene Targets (N, ORF1ab, E, S) | Quantitative detection and variant screening. | Multiplex assays targeting 2-3 genes increase diagnostic confidence. |
| Synthetic RNA Controls (Whole Genome or Fragment) | Quantification standard and extraction/PCR efficiency control. | Must be spiked into sample lysis buffer to monitor full workflow. |
| Buffer EL (Elution Buffer) | Efficient elution of nucleic acids from swab matrices. | Pre-heating to 65°C increases final eluate concentration. |
This technical guide is framed within a comprehensive thesis on SARS-CoV-2 genome structure and its implications for diagnostic target selection. The LoD of molecular diagnostic assays, particularly RT-qPCR, is a critical parameter defining the earliest stage of infection at which the virus can be reliably detected. Optimization hinges on two interdependent pillars: the biochemical composition of the reaction mixture (reagent formulation) and the amplification protocol (cycle optimization). This document provides an in-depth analysis of current strategies to push LoD boundaries for SARS-CoV-2 detection, focusing on the interplay between conserved genomic targets, reagent chemistry, and thermal cycling parameters.
SARS-CoV-2 Genomic Targets and Their Impact on LoD
The selection of amplification targets within the ~30kb SARS-CoV-2 positive-sense single-stranded RNA genome is foundational to assay sensitivity. Conserved regions with minimal variability are prioritized to ensure robustness against emerging variants. The most common targets include:
- Envelope (E) gene: Highly conserved, with a low copy number required for detection, making it a sensitive screening target.
- Nucleocapsid (N) gene: Highly expressed, offering abundant template RNA, but with slightly higher sequence variability.
- RNA-dependent RNA polymerase (RdRp) gene: Part of the ORF1ab region, highly conserved but structurally complex, which can impact primer/probe binding efficiency.
- Spike (S) gene: Useful for variant discrimination but has higher mutation rates, potentially compromising LoD if not carefully managed in multiplex assays.
Optimal reagent formulation must account for the secondary structure and GC content of these specific target regions to ensure efficient reverse transcription and primer annealing.
Reagent Formulation Optimization for Enhanced LoD
The core reaction mix components directly influence enzymatic efficiency, specificity, and the ability to amplify low-copy-number targets.
Key Components and Their Functions
| Component | Function in Enhancing LoD | Optimal Concentration Range (Typical) | Rationale for Optimization |
|---|---|---|---|
| Reverse Transcriptase | Converts viral RNA to cDNA. Processivity and thermostability are critical. | 5-15 U/µL | Use of engineered enzymes with high fidelity and resistance to inhibitors (e.g., heparin, heme) found in clinical samples. |
| DNA Polymerase | Amplifies cDNA. Must have high processivity and strand displacement activity. | 0.5-2 U/µL | Blends of hot-start and proofreading enzymes can improve yield and specificity from low-template reactions. |
| Primers & Probes | Sequence-specific binding for amplification and detection. | 100-900 nM (primers), 50-250 nM (probe) | Chemistries like locked nucleic acid (LNA) or minor groove binder (MGB) increase Tm and specificity, allowing shorter, more efficient probes for challenging targets. |
| MgCl₂ | Essential cofactor for polymerase activity. Influences enzyme fidelity and primer annealing. | 2-6 mM | Titration is crucial; excess Mg²⁺ can reduce specificity, while insufficient amounts lower yield. |
| dNTPs | Building blocks for cDNA synthesis and amplification. | 200-600 µM each | Balanced equimolar mixes are vital. Uracil-DNA glycosylase (UDG) can be added to prevent carryover contamination. |
| Stabilizers/Additives | Protect enzyme activity, reduce nonspecific binding, and mitigate inhibitors. | Varies | Bovine serum albumin (BSA, 0.1-0.5 µg/µL), betaine (0.5-2 M), trehalose (0.2-0.6 M), and non-ionic detergents (e.g., Tween-20) improve reaction robustness. |
Advanced Formulation Strategies
- Surfactant Engineering: Specific non-ionic surfactants (e.g., ThermoPol) can stabilize polymerase at high temperatures, improving performance in the presence of sample-derived inhibitors.
- Cation Optimization: Combining Mg²⁺ with K⁺ and (NH₄)⁺ can optimize polymerase kinetics and primer-template binding stability for specific target sequences.
- pH Buffering Systems: Robust buffering at pH ~8.3 is essential, as the pH shift during amplification can inhibit enzymatic activity in low-template reactions.
Amplification Cycle Optimization
Cycle parameters must be tailored to the reagent formulation to maximize product yield without increasing nonspecific background.
Critical Thermal Cycling Parameters
| Parameter | Standard Range | Optimization for Low-Target Copy Number | Rationale |
|---|---|---|---|
| Reverse Transcription | 50-55°C for 10-30 min | Increase to 55-60°C for 15-20 min | Higher temperature reduces RNA secondary structure, improving processivity for low-copy templates. |
| Initial Denaturation | 95°C for 30 sec - 2 min | 95°C for 2-3 min | Ensures complete separation of RNA:cDNA hybrids and full polymerase activation. |
| Denaturation | 95°C for 5-15 sec | 95°C for 5-10 sec | Shorter time preserves polymerase activity over more cycles. |
| Annealing/Extension | 55-60°C for 30-60 sec | Two-Step: 60-62°C for 30-45 sec.Three-Step: 55-58°C (anneal) & 68-72°C (extend). | A combined, higher-temperature step increases speed and efficiency. For GC-rich targets, a three-step protocol may improve specificity. |
| Cycle Number | 40-45 cycles | Increase to 45-50 cycles | Directly increases probability of detecting low-copy targets. Must be balanced against increased background and reagent depletion. |
Protocol for LoD Determination via Probit Analysis
Objective: To statistically determine the lowest concentration of SARS-CoV-2 target detectable with ≥95% probability. Materials:
- Synthetic RNA control (e.g., from BEI Resources) spanning the E, N, and RdRp genes.
- Optimized master mix (as per formulations above).
- RT-qPCR instrument. Method:
- Prepare a 10-fold serial dilution of the SARS-CoV-2 RNA control in nuclease-free water or negative matrix (e.g., saliva extract). Range: 10⁶ to 10⁰ copies/µL.
- For each dilution level, run a minimum of 20 replicates using the optimized reagent formulation and cycling protocol (50 cycles).
- Record the number of positive replicates (Cq value below a predefined threshold) for each concentration.
- Input the data (log10 concentration vs. proportion of positives) into statistical software (e.g., R, SPSS) and perform probit regression analysis.
- The LoD is defined as the concentration at which the model predicts 95% positivity.
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function/Application | Example Product(s) |
|---|---|---|
| High-Efficiency RT Enzyme | High-temperature, inhibitor-resistant reverse transcription for structured viral RNA. | SuperScript IV, LunaScript RT |
| Hot-Start Polymerase Blends | Provides high sensitivity and specificity by preventing non-specific amplification at low temperatures. | Luna Universal Probe One-Step Mix, TaqPath 1-Step RT-qPCR Master Mix |
| LNA/MGB Probes | Increases hybridization stringency and Tm, allowing for shorter, more specific probes to conserve sequences. | PrimeTime qPCR Probes |
| Inhibitor-Removal Buffers | Additives to master mix or sample prep buffers to neutralize common PCR inhibitors (heme, heparin, humic acid). | OneStep PCR Inhibitor Removal Kit |
| Synthetic RNA Controls | Quantified, sequence-verified RNA for absolute quantification, standard curves, and LoD studies. | BEI Resources SARS-CoV-2 RNA, Twist Synthetic SARS-CoV-2 RNA Controls |
| Digital PCR Master Mix | For absolute quantification and validation of LoD without standard curves; used as a gold-standard comparator. | ddPCR Supermix for Probes (No dUTP) |
Visualized Workflows and Relationships
Title: Optimized One-Step RT-qPCR Workflow for LoD
Title: Factors Converging to Improve Assay LoD
Title: Statistical LoD Determination via Probit Analysis
Benchmarking Diagnostic Performance: Validation, Comparative Analysis, and Regulatory Pathways
Within the broader study of SARS-CoV-2 genome structure and protein targets for diagnostics, assay validation is paramount. The ~30 kb positive-sense RNA genome encodes structural proteins (Spike, Envelope, Membrane, Nucleocapsid) and non-structural proteins (e.g., RdRp, helicase) that serve as primary targets for molecular and antigen tests. This whitepaper details the technical framework for validating novel diagnostic assays against the established gold standards of reverse transcription quantitative PCR (RT-qPCR) and Next-Generation Sequencing (NGS).
Gold Standard Assays: Technical Specifications
RT-qPCR as the Diagnostic Benchmark
RT-qPCR remains the undisputed gold standard for the direct detection of SARS-CoV-2 RNA due to its high sensitivity, specificity, and throughput.
Key SARS-CoV-2 RT-qPCR Targets:
| Genomic Target | Gene/Region | Function/Rationale for Targeting | Assay Examples (WHO-EUL Listed) |
|---|---|---|---|
| Envelope (E) | Structural gene | Highly conserved region; excellent analytical sensitivity. | Charité Berlin assay |
| Nucleocapsid (N) | Structural gene (N1, N2) | Abundant transcript; high copy number improves sensitivity. | CDC (USA) assay |
| RNA-dependent RNA Polymerase (RdRp) | Non-structural gene (nsp12) | Highly specific to SARS-CoV-2; critical for viral replication. | HKU, Pasteur Institute assays |
| Open Reading Frame 1ab (ORF1ab) | Non-structural polyprotein | Large, conserved region; specific to SARS-CoV-2. | China CDC assay |
Critical Performance Metrics for RT-qPCR:
| Metric | Typical Gold Standard Benchmark | Validation Requirement |
|---|---|---|
| Analytical Sensitivity (Limit of Detection) | ≤ 1000 copies/mL (or ≤ 3.5-6.5 genome copies/reaction) | 95% detection rate at LoD. |
| Analytical Specificity | 100% (no cross-reactivity with common respiratory flora/viruses) | In-silico & wet-lab testing against panel. |
| Diagnostic Sensitivity | ≥ 95% (vs. a composite clinical reference) | Testing on confirmed positive clinical samples. |
| Diagnostic Specificity | ≥ 99% (vs. a composite clinical reference) | Testing on confirmed negative/pre-pandemic samples. |
Sequencing as the Confirmatory and Variant-Tracking Benchmark
NGS and Sanger sequencing serve as gold standards for determining viral genome sequence, confirming amplicon specificity, and identifying variants of concern (VOCs).
Sequencing Applications in Validation:
| Application | Platform | Primary Role in Validation |
|---|---|---|
| Amplicon Confirmation | Sanger Sequencing | Verifies the exact nucleotide sequence of RT-qPCR or new assay amplicons. |
| Variant Analysis & Degeneracy Design | Next-Generation Sequencing (NGS) | Identifies mutations in primer/probe binding sites; informs pan-variant assay design. |
| Whole Genome Sequencing (WGS) | NGS (Illumina, Oxford Nanopore) | The ultimate comparator for comprehensive genetic characterization and novel target discovery. |
Experimental Protocols for Comparative Validation
Protocol: Head-to-Head Comparison of New Assay vs. RT-qPCR
Objective: Determine the diagnostic sensitivity and specificity of a new assay (e.g., RT-LAMP, CRISPR-based, antigen test) against RT-qPCR. Materials: See "The Scientist's Toolkit" below. Procedure:
- Sample Panel Curation: Assemble a blinded panel of clinical specimens (e.g., nasopharyngeal swabs in VTM) characterized by a reference RT-qPCR assay. Include a range of Ct values (e.g., 15-35), confirmed negatives, and potential cross-reactants.
- Parallel Testing: Perform the new assay on all samples according to its optimized protocol. Run the reference RT-qPCR assay in parallel on the same sample aliquots.
- Data Analysis:
- Construct a 2x2 contingency table.
- Calculate Sensitivity: (True Positives / (True Positives + False Negatives)) * 100.
- Calculate Specificity: (True Negatives / (True Negatives + False Positives)) * 100).
- Calculate Cohen's Kappa statistic for agreement beyond chance.
Protocol: Sequencing-Based Confirmation of Discrepant Results
Objective: Resolve discordant results between the new assay and RT-qPCR. Procedure:
- Identify Discrepants: Flag all samples with discordant results (e.g., New Assay+/RT-qPCR- and New Assay-/RT-qPCR+).
- Nucleic Acid Extraction: Re-extract RNA from the original specimen.
- Alternative Target RT-qPCR: Test the RNA with a second, orthogonal RT-qPCR assay targeting a different genomic region.
- Sequencing Analysis:
- For New Assay+/RT-qPCR- samples: Perform Sanger sequencing of the new assay's amplicon to confirm it is SARS-CoV-2 specific.
- For New Assay-/RT-qPCR+ samples: Perform Sanger or NGS on the RT-qPCR amplicon to confirm sequence and check for mutations in the new assay's target site.
- Final Reconciliation: Assign true positive/negative status based on composite results from reference RT-qPCR, orthogonal RT-qPCR, and sequencing.
Visualizations
Diagram 1 Title: Validation workflow for new SARS-CoV-2 tests.
Diagram 2 Title: SARS-CoV-2 genome structure and diagnostic targets.
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Function in Validation | Example/Notes |
|---|---|---|
| SARS-CoV-2 Pseudovirus | Safe surrogate for live virus in initial LoD/ specificity studies. | Lentiviral or VSV-based particles expressing SARS-CoV-2 S protein. |
| Inactivated SARS-CoV-2 Virus | Provides native viral RNA & antigen structure for analytical testing. | Gamma-irradiated or chemically inactivated virus (e.g., Delta, Omicron variants). |
| Synthetic RNA Controls | Quantified standards for precise LoD determination and standard curve generation. | Full-length or target-specific RNA transcripts (e.g., from Twist Bioscience). |
| Clinical Sample Panels | Provides real-world matrix for diagnostic performance evaluation. | Well-characterized, remnant de-identified specimens (commercially available). |
| Cross-Reactivity Panels | Validates assay specificity against common pathogens. | Includes nucleic acid/whole organisms of other coronaviruses, influenza, RSV, etc. |
| Human Genomic DNA | Assesses potential for false positives from human background. | Used in specificity panels for assays targeting viral sequences. |
| Master Mixes with UDG/UNG | Prevents amplicon contamination in PCR-based assays. | Contains uracil-DNA glycosylase to degrade carryover amplicons. |
| NGS Library Prep Kits | Prepares samples for sequencing-based confirmation and variant analysis. | Amplicon-based (e.g., Illumina COVIDSeq) or hybridization-capture kits. |
This whitepaper provides an in-depth technical analysis of the clinical performance of diagnostic assays targeting specific SARS-CoV-2 proteins. Within the broader thesis of SARS-CoV-2 genome structure and its implications for diagnostics, we evaluate key metrics—Sensitivity, Specificity, Positive Predictive Value (PPV), and Negative Predictive Value (NPV)—for assays detecting Nucleocapsid (N), Spike (S), and Envelope (E) proteins. Accurate measurement of these metrics is fundamental for the development of effective point-of-care tests, surveillance tools, and therapeutic monitoring assays in both clinical and research settings.
The SARS-CoV-2 genome encodes four primary structural proteins: Spike (S), Nucleocapsid (N), Membrane (M), and Envelope (E). Diagnostic assays predominantly target the N and S proteins due to their abundance and immunogenicity. The N protein is highly expressed during infection and is a major target for antigen-detecting rapid diagnostic tests (Ag-RDTs) and serological assays. The S protein, responsible for viral entry via the ACE2 receptor, is the primary target for neutralizing antibody assays and vaccine efficacy studies. The E and M proteins are less commonly targeted but provide alternative epitopes for assay design, potentially aiding in the detection of variants with mutations in the primary targets.
Core Definitions of Clinical Performance Metrics
- Sensitivity (True Positive Rate): The proportion of actual positive cases correctly identified by the assay. High sensitivity is critical for rule-out tests to minimize false negatives.
- Specificity (True Negative Rate): The proportion of actual negative cases correctly identified by the assay. High specificity is critical for rule-in tests to minimize false positives.
- Positive Predictive Value (PPV): The probability that a subject with a positive test result truly has the condition. PPV is highly dependent on disease prevalence.
- Negative Predictive Value (NPV): The probability that a subject with a negative test result truly does not have the condition. NPV is also prevalence-dependent.
Quantitative Performance Data Across Target Proteins
The following tables summarize recent performance data for diagnostic modalities targeting key SARS-CoV-2 proteins. Data is synthesized from evaluations by regulatory agencies and peer-reviewed clinical studies.
Table 1: Performance of Antigen-Detecting Rapid Diagnostic Tests (Ag-RDTs) by Target Protein
| Target Protein | Assay Format | Avg. Sensitivity (95% CI) | Avg. Specificity (95% CI) | Key Limitations |
|---|---|---|---|---|
| Nucleocapsid (N) | Lateral Flow Immunoassay | 73.0% (68.8-76.9%) | 99.1% (98.8-99.3%) | Sensitivity drops significantly at Ct values >25 (low viral load). |
| Spike (S) | Electrochemiluminescence | 85.2% (79.7-89.5%)* | 98.5% (97.1-99.3%)* | Less abundant than N protein in nasopharyngeal samples. |
| Nucleocapsid (N) | Microfluidic Immunoassay | 93.5% (88.9-96.5%) | 100% (98.8-100%) | Requires specialized reader instrumentation. |
Note: Data for S-targeting Ag-RDTs are limited; this represents a high-complexity laboratory-based antigen assay.
Table 2: Performance of Serological (Antibody) Assays by Target Protein
| Target Protein | Isotype Detected | Avg. Sensitivity >14 days post-onset | Avg. Specificity | Cross-Reactivity Concerns |
|---|---|---|---|---|
| Spike (S1/RBD) | IgG | 98.2% (96.5-99.1%) | 99.5% (98.7-99.8%) | Low; specific to SARS-CoV-2. |
| Nucleocapsid (N) | IgG | 96.5% (94.2-98.0%) | 98.1% (96.2-99.2%) | High with other human coronaviruses (e.g., HCoV-229E, OC43). |
| Spike (S2) | IgG | 90.1% (86.3-93.0%) | 99.0% (97.5-99.6%) | Moderate; conserved across sarbecoviruses. |
Detailed Experimental Protocols
Protocol: Evaluation of an Antigen Rapid Diagnostic Test (Ag-RDT)
Objective: To determine the clinical sensitivity and specificity of a lateral flow assay targeting the SARS-CoV-2 Nucleocapsid (N) protein.
Materials: Suspected patient nasopharyngeal swab samples in viral transport media (VTM), commercially available N-protein Ag-RDT kit, PCR equipment as reference standard, timer.
Procedure:
- Sample Preparation: Vortex VTM samples for 10 seconds. For viscous samples, add 2-3 drops of provided buffer to the extraction tube first.
- Test Execution: Using the transfer pipette, add 3-4 drops of the prepared sample to the sample well (S) of the test cassette.
- Incubation: Start the timer. Allow the test to develop at room temperature (15-30°C) for exactly 15 minutes.
- Result Interpretation: Read results at 15 minutes. Do not interpret after 20 minutes. A visible test line (T) in addition to the control line (C) indicates a positive result. Only a control line indicates a negative result. No control line indicates an invalid test.
- Reference Testing: All samples are tested in parallel via RT-PCR targeting at least two SARS-CoV-2 genes (e.g., N and RdRp).
- Analysis: Calculate Sensitivity = (True Positives / (True Positives + False Negatives)) * 100. Calculate Specificity = (True Negatives / (True Negatives + False Positives)) * 100.
Protocol: Neutralizing Antibody Assay using Pseudotyped Virus
Objective: To measure the presence and titer of neutralizing antibodies against the SARS-CoV-2 Spike protein in patient serum.
Materials: HEK293T-ACE2 cells, plasmid expressing SARS-CoV-2 Spike protein (variant of interest), lentiviral backbone plasmid (e.g., pNL4-3.Luc.R-E-), patient serum samples, cell culture media, luciferase assay kit.
Procedure:
- Pseudovirus Production: Co-transfect HEK293T cells with the Spike plasmid and the lentiviral backbone plasmid using a transfection reagent. Harvest pseudotyped virus supernatant at 48 and 72 hours post-transfection.
- Serum Heat-Inactivation: Heat patient serum at 56°C for 30 minutes to inactivate complement.
- Neutralization Assay: In a 96-well plate, serially dilute heat-inactivated serum. Mix each dilution with an equal volume of pseudovirus stock. Incubate at 37°C for 1 hour.
- Infection: Add the serum-virus mixture to a plate of HEK293T-ACE2 cells. Include virus-only (no serum) and cell-only controls.
- Incubation & Measurement: After 48-72 hours, lyse cells and measure luciferase activity. The readout is Relative Luminescence Units (RLU).
- Analysis: Calculate % neutralization = [1 - (RLU of serum sample / RLU of virus-only control)] * 100. The neutralization titer (NT50) is the serum dilution that inhibits 50% of the luciferase signal, calculated using non-linear regression.
Visualizations
Title: Ag-RDT Validation Workflow Against PCR Reference
Title: Spike Protein Neutralization Pathway
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for SARS-CoV-2 Protein-Targeted Diagnostics Research
| Reagent / Material | Primary Function in Research/Assay | Key Consideration |
|---|---|---|
| Recombinant SARS-CoV-2 Proteins (N, S1, RBD, E) | Used as calibration standards, for assay development, and for antibody characterization. | Source from vendors providing full sequence and post-translational modification data. Critical for variant-specific studies. |
| Monoclonal Antibodies (Anti-N, Anti-S) | Capture/detection pairs for immunoassay development. Neutralizing mAbs for control materials. | Define epitope mapping. Validate for lack of cross-reactivity with common human coronavirus proteins. |
| Pseudotyped Virus Systems (Lentiviral/VSV) | Safe, BSL-2 surrogate for live virus in neutralization assays. Can incorporate Spike variants. | Verify packaging efficiency and single-round infectivity. Match pseudovirus backbone to detection readout (luciferase, GFP). |
| ACE2 Expressing Cell Lines | Essential substrate for Spike-mediated entry and neutralization assays. | Monitor ACE2 expression stability across passages. HEK293T-ACE2 is standard; Caco-2 cells offer endogenous expression. |
| Serum/Plasma Panels (Convalescent, Pre-pandemic) | Gold-standard positive/negative controls for serology and test validation. | Ensure panels are well-characterized (PCR-confirmed, symptom/days post-onset data included). Pre-pandemic panels assess specificity. |
| Synthetic SARS-CoV-2 RNA | Positive control for molecular assay development, unaffected by supply chain issues for live virus. | Contains full-length target sequences (e.g., N gene) with defined copy number for sensitivity (LOD) studies. |
This analysis is framed within a broader thesis examining the SARS-CoV-2 genome structure to identify optimal protein targets for diagnostic applications. The viral genome encodes four major structural proteins: Spike (S), Envelope (E), Membrane (M), and Nucleocapsid (N). For antigen-detecting rapid diagnostic tests (Ag-RDTs), the S and N proteins emerge as primary candidates due to their abundance and distinct biological roles. This whitepaper provides a comparative technical analysis, weighing the high immunogenicity of the S-protein against the superior stability of the N-protein as core considerations for assay design.
Biological & Structural Fundamentals
Spike (S) Protein: A class I fusion trimer protruding from the viral envelope. It comprises two subunits: S1 (containing the Receptor-Binding Domain, RBD) for ACE2 attachment and S2 for membrane fusion. Its high surface accessibility and immunodominance drive strong antibody responses.
Nucleocapsid (N) Protein: An RNA-binding protein that complexes with the viral genomic RNA to form the ribonucleoprotein (RNP) core. It is involved in genome packaging, virion assembly, and modulating host cell processes. It is abundantly expressed inside the virion and infected cells.
Quantitative Comparison: Stability vs. Immunogenicity
Table 1: Core Comparison of S-Protein and N-Protein as Antigen Test Targets
| Parameter | Spike (S) Protein | Nucleocapsid (N) Protein |
|---|---|---|
| Primary Localization | Viral envelope surface | Viral core & infected cell cytoplasm |
| Copy Number per Virion | ~24-40 trimers | ~1000 molecules |
| Immunogenicity | Extremely high; primary target for neutralizing antibodies | High; dominant target for T-cell & non-neutralizing Ab responses |
| Genetic Stability | Lower; RBD is a hotspot for mutations (e.g., Omicron variants) | Higher; more conserved sequence, though variations occur |
| Epitope Stability | Conformational epitopes susceptible to disruption | Linear epitopes largely resistant to denaturation |
| Expression in Infected Cells | Moderate, primarily surface/secreted | Very high, cytoplasmic accumulation |
| Impact of Vaccination | High anti-S Ab from vaccination can obscure infection signal | Minimal; most vaccines are S-based, leaving anti-N Ab as infection marker |
Table 2: Performance Metrics in Antigen Tests (Representative Data)
| Metric | S-Targeting Tests | N-Targeting Tests |
|---|---|---|
| Analytical Sensitivity (LoD) | ~10^2-10^3 TCID50/mL | ~10^1-10^2 TCID50/mL |
| Clinical Sensitivity (vs. PCR)* | ~84% (75-92%) | ~94% (88-98%) |
| Specificity | >98% | >99% |
| Heat Stability (e.g., 37°C for 4 wks) | Moderate signal decay | High signal retention |
| Variant Reactivity (e.g., Omicron) | Potentially reduced due to mutations | Largely maintained |
*Data aggregated from multiple clinical evaluations; sensitivity varies with viral load and symptom status.
Detailed Experimental Protocols
Protocol 4.1: Comparative Thermostability Assay for Antigen Test Strips Objective: To quantify the degradation of immobilized S and N antigens on lateral flow strips under accelerated storage conditions. Materials: See "Scientist's Toolkit" (Section 6). Method:
- Strip Preparation: Coat control lines with anti-species IgG. For test lines, use nitrocellulose strips to immobilize identical molar quantities of recombinant S-trimer (stabilized) and N-protein.
- Accelerated Aging: Store triplicate sets of strips in sealed desiccant pouches at 37°C, 45°C, and 4°C (control) for 0, 2, 4, and 8 weeks.
- Signal Challenge: At each time point, run strips with a standardized positive control (heat-inactivated SARS-CoV-2 lysate in universal transport medium) and negative control.
- Quantification: Use a lateral flow reader to measure test line intensity (pixel density or RFU).
- Analysis: Plot signal intensity vs. time for each temperature. Calculate the decay constant (k) and half-life of the antigen activity on the strip.
Protocol 4.2: Epitope Mapping for Variant Cross-Reactivity Objective: To assess the impact of S and N protein mutations on monoclonal antibody (mAb) binding used in test kits. Method:
- Protein Array: Spot recombinant wild-type and variant (e.g., Omicron BA.5, XBB.1.5) S-RBD and full-length N-protein onto nitrocellulose slides.
- Antibody Probing: Incubate arrays with HRP-conjugated mAbs (at identical concentrations) used in commercial Ag-RDTs (e.g., S-targeting mAb #1, N-targeting mAb #2).
- Detection: Develop with chemiluminescent substrate and image with a microarray scanner.
- Data Normalization: Express variant spot intensity as a percentage of wild-type intensity. A drop below 70% indicates significant epitope disruption.
Visualization Diagrams
Diagram Title: Decision Pathway for Selecting S or N Protein Target
Diagram Title: Dual-Target S & N Protein Lateral Flow Assay Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Antigen Target Validation
| Reagent / Material | Function & Rationale |
|---|---|
| Recombinant S-trimer (HexaPro stabilized) | Mimics native conformation for assessing conformational antibody binding and test development. |
| Recombinant full-length N-protein | Source of linear epitopes for monoclonal antibody characterization and as a calibrator. |
| HRP-conjugated anti-N mAb panel | Critical for epitope binning and mapping studies to identify non-competing pairs for sandwich assays. |
| Pseudotyped VSV particles (with variant S) | Safe, BSL-2 surrogate for live virus to evaluate test performance against variants of concern. |
| Clinical specimen panel (heat-inactivated) | Well-characterized PCR-positive/negative samples for clinical sensitivity/specificity studies. |
| Lateral flow nitrocellulose cards & reader | Platform for prototype strip development and objective, quantitative signal measurement. |
| Accelerated stability chamber | To simulate long-term storage conditions (elevated T & humidity) for product shelf-life studies. |
The development of diagnostic assays for SARS-CoV-2 relies on a precise understanding of its genome structure and protein targets. The ~30 kb positive-sense RNA genome encodes four structural proteins (Spike (S), Envelope (E), Membrane (M), Nucleocapsid (N)) and multiple non-structural and accessory proteins. The N and S proteins are primary targets for antigen and molecular assays, respectively. Regulatory pathways for these assays, including FDA Emergency Use Authorization (EUA), WHO Emergency Use Listing (EUL), and CE-IVD marking, demand rigorous validation against these genomic targets to ensure clinical accuracy and public health utility.
Core Regulatory Pathways: Comparative Analysis
FDA Emergency Use Authorization (EUA)
Governing Body: U.S. Food and Drug Administration (FDA) Legal Basis: Section 564 of the Federal Food, Drug, and Cosmetic Act Primary Purpose: To allow the use of unapproved medical products during a declared public health emergency.
Key Technical Requirements for SARS-CoV-2 Assays:
- Analytical Sensitivity (LoD): Must be established using genetically diverse clinical isolates or synthetic targets covering all known variants. The LoD must be verified in the matrix of intended use (e.g., nasal swabs, saliva).
- Analytical Specificity: Testing must include cross-reactivity with common respiratory pathogens (e.g., influenza, RSV, endemic coronaviruses) and human genomic DNA.
- Clinical Evaluation: For molecular assays, positive percent agreement (PPA) and negative percent agreement (NPA) versus an authorized comparator assay are required. PPA is critical for symptomatic populations.
- Quality Controls: Must include internal and external controls to monitor extraction, amplification, and detection.
WHO Emergency Use Listing (EUL)
Governing Body: World Health Organization (WHO) Purpose: To expedite the availability of essential health products during a Public Health Emergency of International Concern (PHEIC) for procurement and use globally, especially in low- and middle-income countries.
Key Technical Requirements:
- Performance Data: Requires data comparable to FDA EUA, with emphasis on inclusivity across global SARS-CoV-2 variants.
- Risk-Benefit Analysis: A comprehensive assessment considering the public health need and the assay's performance.
- Manufacturing Quality: Compliance with international standards for quality management systems (e.g., ISO 13485).
- Stability & Labeling: Demonstrated stability under expected supply chain conditions and labeling suitable for global use.
CE-IVD Marking
Governing Body: Notified Bodies under the European Union's In Vitro Diagnostic Regulation (IVDR) 2017/746. Legal Basis: IVDR is fully applicable as of May 2022. Purpose: To certify that an in vitro diagnostic device complies with EU health, safety, and performance requirements for placement on the European market.
Key Technical Requirements under IVDR:
- Performance Evaluation Report (PER): A comprehensive document including scientific validity, analytical performance, and clinical performance data.
- General Safety and Performance Requirements (GSPR): Adherence to all applicable Annex I requirements of the IVDR.
- Risk Management: Implementation of a systematic risk management process per ISO 14971.
- Post-Market Surveillance (PMS): Establishment of a proactive PMS plan and a Post-Market Performance Follow-up (PMPF) plan.
Comparative Data Tables
Table 1: Key Performance and Submission Requirements
| Criterion | FDA EUA | WHO EUL | CE-IVD (Under IVDR) |
|---|---|---|---|
| Legal Framework | FD&C Act, Section 564 | WHO Emergency Use Procedure | Regulation (EU) 2017/746 (IVDR) |
| Validity Period | Duration of the declared emergency | Duration of the PHEIC; subject to reassessment | 5-year certificate (typical) |
| Lead Time (Typical) | 2-8 weeks (expedited review) | 2-12 weeks | 12-18+ months (full conformity assessment) |
| Clinical Evidence | PPA/NPA vs. comparator (n≥30 positive, n≥30 negative samples minimum) | Comparable performance data; often accepts data from other stringent reviews | Clinical performance study per IVDR Annex XIII, considering intended purpose and population |
| Quality System | Expectation of compliant QMS (e.g., 21 CFR Part 820) | ISO 13485 or equivalent | ISO 13485 mandatory for Class C/D devices (most SARS-CoV-2 assays) |
| Post-Market Obligations | Adverse event reporting, periodic updates to FDA | Reporting of performance issues, updates to WHO | Active PMS plan, PMPF studies, Periodic Safety Update Report (PSUR) |
Table 2: Analytical Validation Benchmarks for a SARS-CoV-2 qPCR Assay
| Validation Parameter | Typical Requirement | Relevant SARS-CoV-2 Genomic Target |
|---|---|---|
| Limit of Detection (LoD) | ≤ 1000 genome copies/mL (or copies/swab) | Confirm with at least two gene targets (e.g., N, E, RdRp) |
| Inclusivity (Variant Detection) | Must detect all known circulating variants of concern (Alpha, Delta, Omicron, etc.) | Testing with in silico analysis and wet lab testing on synthetic or clinical variant samples |
| Cross-Reactivity (Analytical Specificity) | No cross-reactivity with ≥ 30 common respiratory pathogens and human microbiomes | Wet lab testing against a panel of pathogens; in silico BLAST against human genome and microbiome |
| Precision (Repeatability & Reproducibility) | ≥ 95% agreement across operators, days, and instrument lots | Testing at 2-3 concentrations (near LoD, mid, high) across the intended sample matrix |
Experimental Protocols for Assay Validation
Protocol 1: Determination of Limit of Detection (LoD) for a SARS-CoV-2 RT-qPCR Assay Objective: To establish the lowest concentration of SARS-CoV-2 viral particles at which the assay detects ≥95% of replicates.
Materials:
- SARS-CoV-2 genomic RNA standard (quantified, full-length or target segment).
- Negative clinical matrix (e.g., pooled nasal swab transport media).
- RT-qPCR master mix, primers/probes targeting N1 and RdRp genes.
- Real-Time PCR system.
Methodology:
- Serial Dilution: Prepare a dilution series of the SARS-CoV-2 RNA standard in the negative clinical matrix. A typical range is from 10^5 to 10^0 copies/µL.
- Replicate Testing: Test each dilution level in a minimum of 20 replicates across multiple runs/days.
- Data Analysis: Calculate the detection rate at each concentration. Use probit or logit regression analysis to determine the concentration at which 95% of replicates are detected. This is the provisional LoD.
- Verification: Test the provisional LoD in 20 independent replicates. The assay must detect at least 19/20 (95%) to verify the LoD.
Protocol 2: Inclusivity (Variant) Testing via In Silico Analysis Objective: To computationally assess if primer/probe sequences bind effectively to all known SARS-CoV-2 variants.
Materials:
- Primer and probe sequences (FASTA format).
- Database of SARS-CoV-2 variant genome sequences (e.g., GISAID, NCBI Virus).
- Sequence alignment software (e.g., Geneious, CLC Workbench, BLAST+).
Methodology:
- Sequence Retrieval: Download a representative set of genome sequences for each Variant of Concern (VOC) and Variant of Interest (VOI).
- Multiple Sequence Alignment: Align all variant sequences with the reference Wuhan-Hu-1 strain (NC_045512.2).
- Target Region Extraction: Extract the genomic regions corresponding to your assay's primer and probe binding sites from the aligned dataset.
- Mismatch Analysis: Manually or algorithmically identify nucleotide mismatches within the last 5 bases of the 3' end of primers (critical for extension) and within the probe sequence.
- Impact Prediction: Report variants with mismatches. Design and test alternative primers/probes if critical mismatches are predicted to impact sensitivity.
Visualizations
FDA EUA Review Workflow
IVDR Device Classification Logic
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for SARS-CoV-2 Diagnostic Assay Development
| Reagent / Material | Function / Role in Development | Example/Target |
|---|---|---|
| Quantified SARS-CoV-2 RNA Standards | Serves as positive control for LoD determination, calibration, and precision studies. Must represent full genome or key targets (N, S, RdRp). | Genomic RNA from cultured virus (heat-inactivated) or synthetic controls (e.g., from Twist Bioscience, ATCC). |
| Inactivated SARS-CoV-2 Virus | Used for specificity testing, interference studies, and safety training. Provides native virion structure. | Gamma-irradiated or chemically inactivated virus (e.g., ZeptoMetrix, The Native Antigen Company). |
| Pseudotyped Virus Particles | Safe alternative for neutralization studies or assay development targeting the Spike protein entry mechanism. | Lentiviral particles pseudotyped with SARS-CoV-2 S protein. |
| Recombinant Viral Proteins | Critical for developing immunoassays (e.g., lateral flow). Used as capture/detection antigens and for antibody specificity testing. | Recombinant N, S (S1, RBD), M, E proteins (e.g., Sino Biological, Acro Biosystems). |
| Clinical Sample Panels | For clinical validation. Must be well-characterized (positive/negative status, CT values, variant typing). | Panels from commercial biorepositories (e.g., Discovery Life Sciences) or institutional IRB-approved collections. |
| Pathogen Genomic DNA/RNA Panels | For analytical specificity (cross-reactivity) testing. Includes common respiratory flora and viruses. | Panels from ATCC or ZeptoMetrix. |
| Human Biological Matrices | For interference testing and as a diluent for standards. Includes saliva, nasal swab media, blood, etc. | Purchased from commercial biorepositories or pooled from donors under IRB protocol. |
Within the broader thesis on SARS-CoV-2 genome structure and its implications for diagnostics research, the non-structural proteins (Nsps) encoded by the replicase polyprotein (ORF1a/b) present a high-value, yet underexploited, target landscape. While nucleocapsid (N) and spike (S) proteins dominate current diagnostic and therapeutic efforts, Nsps offer distinct advantages for future preparedness: they are highly conserved across sarbecoviruses, essential for viral replication, and less prone to mutation under host immune pressure. This technical guide evaluates emerging Nsp targets and details the integration of multiplex detection platforms, establishing a paradigm for proactive diagnostic development against future viral threats.
Evaluation of Novel Non-Structural Protein (Nsp) Targets
Nsp proteins, generated by extensive proteolytic processing of ORF1a/b, form the core replication-transcription complex (RTC). Their conserved nature makes them prime candidates for pan-sarbecovirus detection and broad-spectrum therapeutic intervention.
Key Nsp Targets for Diagnostics:
- Nsp3 (Papain-like protease, PLpro): Involved in immune evasion by cleaving ISG15 and ubiquitin from host proteins. Its conserved domains are ideal for nucleic acid amplification test (NAAT) design.
- Nsp5 (Main protease, Mpro/3CLpro): Absolutely essential for viral polyprotein processing. Its active site is highly conserved, making it a top target for both antiviral drugs and CRISPR-based diagnostic assays.
- Nsp12 (RNA-dependent RNA polymerase, RdRp): The central enzyme of viral replication. The most conserved region across all RNA viruses, providing a "universal" coronavirus target.
- Nsp13 (Helicase): A conserved enzyme with 5'-to-3' helicase and RNA 5'-triphosphatase activities, crucial for RNA capping and replication.
Quantitative Comparison of Diagnostic Target Regions (2023-2024 Data)
Table 1: Genetic Conservation and Diagnostic Performance of SARS-CoV-2 Protein Targets
| Target Protein | Genomic Region | Average % Identity Across Sarbecoviruses (Range) | Reported LoD for qPCR Assays (copies/µL) | Mutation Rate (Relative to S protein) |
|---|---|---|---|---|
| Spike (S) | ORF2 | ~76% (Variable RBD) | 10-100 | 1.0 (Reference) |
| Nucleocapsid (N) | ORF9 | ~88% | 5-50 | 0.6 |
| Nsp5 (Mpro) | ORF1a | ~96% | 1-10 | 0.2 |
| Nsp12 (RdRp) | ORF1b | ~98% | 1-10 | 0.1 |
| Nsp13 (Helicase) | ORF1b | ~95% | 5-20 | 0.3 |
Multiplex Platform Architectures for Future Preparedness
Multiplexing is critical for differentiating SARS-CoV-2 from other respiratory pathogens and its variants, while also surveilling for novel viruses.
A. High-Throughput Nucleic Acid-Based Platforms:
- Next-Generation Sequencing (NGS): Enables unbiased pan-viral detection and variant characterization.
- Multiplexed CRISPR-Cas Systems: Utilizing Cas12a and Cas13 in tandem for simultaneous detection of multiple conserved targets (e.g., Nsp12 + Nsp5 + human control) on lateral flow readouts.
- Digital PCR (dPCR) Panels: Ultra-sensitive absolute quantification of 5-10 viral targets, including conserved Nsp regions, for precise viral load tracking and early emergence detection.
B. Multiplexed Immunoassay Platforms:
- Luminex xMAP Technology: Bead-based assays to simultaneously detect IgG/IgM responses against a panel of antigens (S, N, Nsp3, Nsp13).
- Multiplexed Electrochemical Biosensors: Array-based sensors with immobilized probes for different Nsp proteins, enabling rapid, point-of-care differential diagnosis.
Experimental Protocols
Protocol 1: Multiplex RT-qPCR for Conserved Nsp Targets (Two-Tube Assay)
- Primer/Probe Design: Design TaqMan probes for Nsp5 (FAM), Nsp12 (HEX), and human RNase P (Cy5) as internal control. Use conserved regions identified via multiple sequence alignment (Clustal Omega).
- RNA Extraction: Use magnetic bead-based kits (e.g., silica membrane) from nasopharyngeal swab in VTM.
- Master Mix Preparation (Per Reaction):
- 5 µL 4x TaqPath 1-Step RT-qPCR Master Mix
- 1 µL Primer/Probe Mix (each at final conc. 400 nM primer, 200 nM probe)
- 2 µL Nuclease-free water
- 2 µL RNA template
- Thermocycling Conditions:
- Reverse Transcription: 25°C for 10 min, 50°C for 15 min.
- PCR Activation: 95°C for 2 min.
- 45 Cycles: Denature at 95°C for 3 sec, Anneal/Extend at 60°C for 30 sec (acquire fluorescence).
Protocol 2: Multiplex Lateral Flow Assay using CRISPR-Cas12a
- RPA Amplification: Perform isothermal amplification (20 min, 42°C) of Nsp5 and Nsp12 targets using specific primers.
- CRISPR Detection: Incubate amplicon with a cocktail containing:
- Cas12a enzyme (2 µM)
- crRNA specific for Nsp5 (labeled with FAM)
- crRNA specific for Nsp12 (labeled with DIG)
- Fluorescent quencher reporters (FQ for FAM, another for DIG)
- Lateral Flow Readout: Apply mixture to a strip with two test lines: anti-FAM (for Nsp5) and anti-DIG (for Nsp12). Cleavage of reporters by activated Cas12a prevents line appearance. Line absence = Positive detection.
Visualization: Pathways and Workflows
Diagram Title: Nsp Protein Role in Diagnostic Targeting
Diagram Title: Multiplex Diagnostic Platform Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Nsp-Targeted Multiplex Diagnostic Research
| Reagent/Material | Function | Example (Brand) |
|---|---|---|
| Consensus Nsp Primers/Probes | For specific amplification/detection of conserved regions in Nsp5, Nsp12, Nsp13. | IDT PrimeTime qPCR Assays |
| Multiplex RT-qPCR Master Mix | Enzymatic mix optimized for simultaneous amplification of >4 targets with minimal cross-talk. | Thermo Fisher TaqPath 1-Step Multiplex Master Mix |
| CRISPR-Cas12a/Cas13 Enzyme Kits | For developing highly specific, amplification-coupled lateral flow or fluorescent assays. | BioVision CRISPR Cas12a Detection Kit |
| Magnetic Bead RNA Extraction Kits | High-throughput, automated nucleic acid purification from diverse sample matrices. | Qiagen QIAamp 96 Virus QIAcube HT Kit |
| Luminex Magnetic Bead Panels | Customizable bead sets for coupling with recombinant Nsp antigens for serology. | Luminex xMAP Antibody Coupling Kits |
| Recombinant Nsp Proteins | Positive controls for assay development, standard curves, and immunization. | Sino Biological SARS-CoV-2 Nsp5, Nsp12, Nsp13 Proteins |
| Digital PCR (dPCR) Chips/Cartridges | Partitioning technology for absolute quantification of multiple viral targets without standards. | Bio-Rad QX200 ddPCR System |
| Lateral Flow Strips (Biotin/Test Line) | Readout platform for multiplex CRISPR or immunoassay results. | Abcam PCRD Lateral Flow Device |
Conclusion
The SARS-CoV-2 genome presents a defined yet evolving set of protein targets that are foundational to modern diagnostic strategies. A successful diagnostic pipeline requires a deep understanding of genomic structure (Intent 1), applied through robust methodological design (Intent 2), which must be continually optimized against viral evolution and technical pitfalls (Intent 3), and rigorously validated through comparative clinical and regulatory analysis (Intent 4). Future directions necessitate agile assay design platforms capable of rapid redeployment, increased reliance on multiplexing to guard against target degradation, and the integration of genomic surveillance directly into diagnostic pipelines. For the research and development community, this underscores the imperative to target highly conserved regions while developing adaptable protein-binding platforms, ensuring diagnostic resilience against future viral variants and pandemics.